Expo
view channel
view channel
view channel
view channel
view channel
view channel
view channel
view channel
view channel
Clinical Chem.Molecular DiagnosticsHematologyImmunologyMicrobiologyPathologyTechnologyIndustry
Events
Webinars

- Simple Dual-Tau Blood Test Detects and Stages Alzheimer’s Disease
- Alzheimer’s Blood Biomarkers Linked to Early Cognitive Differences Before Dementia
- Urine-Based Test Shows Promise for Autism Screening in Children
- Liquid Biopsy Biomarkers May Improve Childhood Epilepsy Diagnosis
- Blood-Based Sensor Detects Early Signs of Alzheimer’s and Parkinson’s
- Circulating Tumor DNA Testing Guides Chemotherapy, Reduces Relapse in Colon Cancer
- Researchers Uncover Distinct Chromosome Signature in Aggresive ALT Cancers
- Genomic Test Guides Chemotherapy Decisions in Early-Stage Breast Cancer
- Simple Cytogenetic Method Could Improve Classification of ALL Subtypes
- Blood-Based Assay Enables Noninvasive Monitoring of Sarcoma Immunotherapy Response
- Blood Eosinophil Count May Predict Cancer Immunotherapy Response and Toxicity
- Higher Ferritin Threshold May Improve Iron Deficiency Detection in Children
- Stem Cell Biomarkers May Guide Precision Treatment in Acute Myeloid Leukemia
- Advanced CBC-Derived Indices Integrated into Hematology Platforms
- Blood Test Enables Early Detection of Multiple Myeloma Relapse
- Study Points to Autoimmune Pathway Behind Long COVID Symptoms
- Metabolic Biomarker Distinguishes Latent from Active Tuberculosis and Tracks Treatment Response
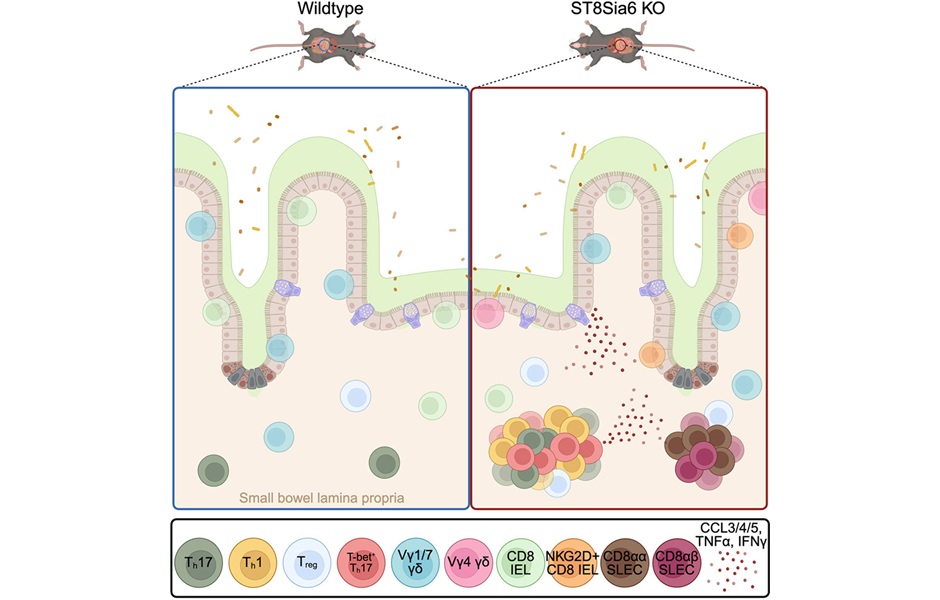
- Immune Enzyme Linked to Treatment-Resistant Inflammatory Bowel Disease
- Simple Blood Test Could Replace Biopsies for Lung Transplant Rejection Monitoring
- Routine TB Screening Test May Reveal Immune Aging and Mortality Risk
- FDA-Cleared Gastrointestinal Panel Detects 24 Pathogen Targets
- New AMR Assay Supports Rapid Infection Control Screening in Hospitals
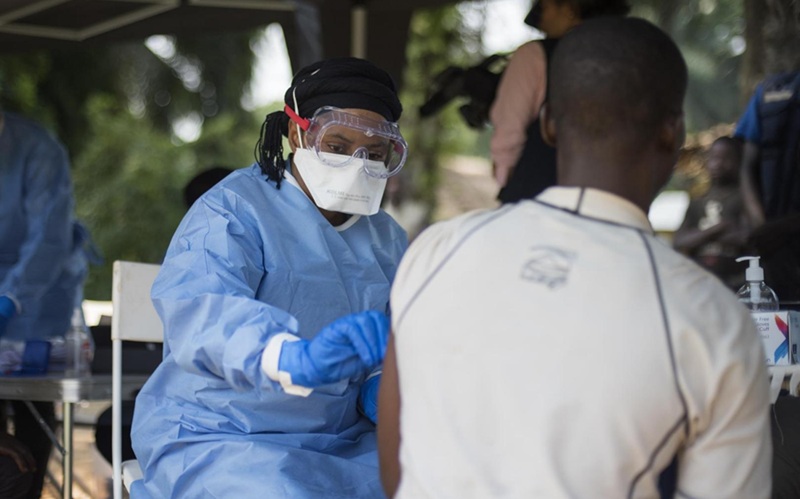
- Diagnostic Gaps Complicate Bundibugyo Ebola Outbreak Response in Congo
- Study Finds Hidden Mpox Infections May Drive Ongoing Spread
- Large-Scale Genomic Surveillance Tracks Resistant Bacteria Across European Hospitals
- Agentic AI Platform Supports Genomic Decision-Making in Oncology
- Mailed Screening Kits Help Reduce Colorectal Cancer Screening Gaps
- Algorithm Panel Aids Liver Fibrosis Assessment and Liver Cancer Surveillance
- AI-Enabled Assistant Unifies Molecular Workflow Planning and Support
- AI Tool Automates Validation of Laboratory Software Configuration Changes
- Partnership Expands Access to Alzheimer’s Blood Tests in Latin America and Caribbean
- Global Multiplex Assays Market Driven by High-Throughput Diagnostic Demand
- Global Framework Integrates Digital Pathology for Companion Diagnostic Development
- GRAIL Presents Full Results from NHS-Galleri Trial at ASCO 2026
- Werfen and Oxford Nanopore Collaborate on Transplant Assay Development
- Tumor Genome Marker May Predict Treatment Benefit in Pediatric Cancers
- Lysosomal Gene Defect Linked to Severe Childhood Brain Disorders
- Genetic Testing Identifies Greater Inherited Sudden Cardiac Arrest Risk in Younger Individuals
- Hidden 'Jumping Gene' Variant Linked to Higher Pancreatic Cancer Risk
- Common White Blood Cells Produce Schizophrenia-Linked Protein
- FDA Approval Expands Automated PD-L1 Testing Across Solid Tumors
- AI-Powered Atlas Maps Immune Structures Linked to Cancer Outcomes
- AI Tool Extracts Immune Signals from Biopsy to Inform Myeloma Therapy
- Rapid AI Tool Predicts Cancer Spatial Gene Expression from Pathology Images
- AI Pathology Test Receives FDA Breakthrough for Bladder Cancer Risk Stratification

 Expo
Expo
- Simple Dual-Tau Blood Test Detects and Stages Alzheimer’s Disease
- Alzheimer’s Blood Biomarkers Linked to Early Cognitive Differences Before Dementia
- Urine-Based Test Shows Promise for Autism Screening in Children
- Liquid Biopsy Biomarkers May Improve Childhood Epilepsy Diagnosis
- Blood-Based Sensor Detects Early Signs of Alzheimer’s and Parkinson’s
- Circulating Tumor DNA Testing Guides Chemotherapy, Reduces Relapse in Colon Cancer
- Researchers Uncover Distinct Chromosome Signature in Aggresive ALT Cancers
- Genomic Test Guides Chemotherapy Decisions in Early-Stage Breast Cancer
- Simple Cytogenetic Method Could Improve Classification of ALL Subtypes
- Blood-Based Assay Enables Noninvasive Monitoring of Sarcoma Immunotherapy Response
- Blood Eosinophil Count May Predict Cancer Immunotherapy Response and Toxicity
- Higher Ferritin Threshold May Improve Iron Deficiency Detection in Children
- Stem Cell Biomarkers May Guide Precision Treatment in Acute Myeloid Leukemia
- Advanced CBC-Derived Indices Integrated into Hematology Platforms
- Blood Test Enables Early Detection of Multiple Myeloma Relapse
- Study Points to Autoimmune Pathway Behind Long COVID Symptoms
- Metabolic Biomarker Distinguishes Latent from Active Tuberculosis and Tracks Treatment Response
- Immune Enzyme Linked to Treatment-Resistant Inflammatory Bowel Disease
- Simple Blood Test Could Replace Biopsies for Lung Transplant Rejection Monitoring
- Routine TB Screening Test May Reveal Immune Aging and Mortality Risk
- FDA-Cleared Gastrointestinal Panel Detects 24 Pathogen Targets
- New AMR Assay Supports Rapid Infection Control Screening in Hospitals
- Diagnostic Gaps Complicate Bundibugyo Ebola Outbreak Response in Congo
- Study Finds Hidden Mpox Infections May Drive Ongoing Spread
- Large-Scale Genomic Surveillance Tracks Resistant Bacteria Across European Hospitals
- Agentic AI Platform Supports Genomic Decision-Making in Oncology
- Mailed Screening Kits Help Reduce Colorectal Cancer Screening Gaps
- Algorithm Panel Aids Liver Fibrosis Assessment and Liver Cancer Surveillance
- AI-Enabled Assistant Unifies Molecular Workflow Planning and Support
- AI Tool Automates Validation of Laboratory Software Configuration Changes
- Partnership Expands Access to Alzheimer’s Blood Tests in Latin America and Caribbean
- Global Multiplex Assays Market Driven by High-Throughput Diagnostic Demand
- Global Framework Integrates Digital Pathology for Companion Diagnostic Development
- GRAIL Presents Full Results from NHS-Galleri Trial at ASCO 2026
- Werfen and Oxford Nanopore Collaborate on Transplant Assay Development
- Tumor Genome Marker May Predict Treatment Benefit in Pediatric Cancers
- Lysosomal Gene Defect Linked to Severe Childhood Brain Disorders
- Genetic Testing Identifies Greater Inherited Sudden Cardiac Arrest Risk in Younger Individuals
- Hidden 'Jumping Gene' Variant Linked to Higher Pancreatic Cancer Risk
- Common White Blood Cells Produce Schizophrenia-Linked Protein
- FDA Approval Expands Automated PD-L1 Testing Across Solid Tumors
- AI-Powered Atlas Maps Immune Structures Linked to Cancer Outcomes
- AI Tool Extracts Immune Signals from Biopsy to Inform Myeloma Therapy
- Rapid AI Tool Predicts Cancer Spatial Gene Expression from Pathology Images
- AI Pathology Test Receives FDA Breakthrough for Bladder Cancer Risk Stratification