Expo
view channel
view channel
view channel
view channel
view channel
view channel
view channel
view channel
view channel
Clinical Chem.
HematologyImmunologyMicrobiologyPathologyTechnologyIndustry
Events
Webinars

- Routine Blood Tests Years Before Pregnancy Could Identify Preeclampsia Risk
- Blood Test Detects Testicular Cancer Missed by Standard Markers
- Routine Blood Tests Identify Biomarkers Linked to PTSD
- Proteomic Data Underscore Need for Age-Specific Pediatric Reference Ranges
- Routine Blood Count Ratio Linked to Future Alzheimer’s and Dementia Risk
- Blood Test Maps Tumor Microenvironment to Predict Immunotherapy Response
- Multiplex Respiratory Panel Integrates Automated Extraction to Streamline High-Volume Testing
- Whole-Blood RNA Test Predicts Disease Trajectory and Treatment Response
- Blood-Based Epigenetic Test Predicts GLP-1 Response and Tracks Treatment Effects
- Tumor Genomic Testing Guides Immunotherapy Selection in Pituitary Tumors
- Blood Test Enables Early Detection of Multiple Myeloma Relapse
- Single Assay Enables Rapid HLA and ABO Genotyping for Transplant Matching
- Prognostic Biomarker Identified in Diffuse Large B-Cell Lymphoma
- Routine Blood Test Parameters Link Anemia to Cancer Risk and Mortality
- Prognostic Tool Guides Personalized Treatment in Rare Blood Cancer
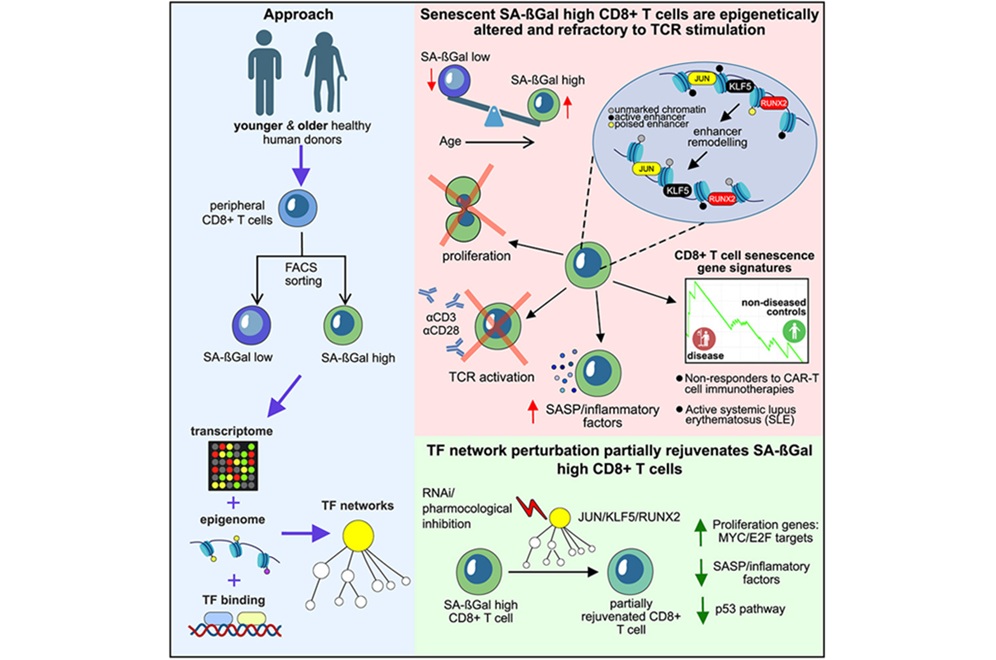
- T-Cell Senescence Profiling May Predict CAR T Responses
- Finger-Prick Lateral Flow Test Detects Sepsis Biomarkers at Point of Care
- Study Highlights Low Sensitivity of Current Lyme Tests in Early Infection
- Immune Aging Clock Quantifies Immunosenescence and Identifies Therapeutic Target
- Study Finds Influenza Often Undiagnosed in Winter Deaths
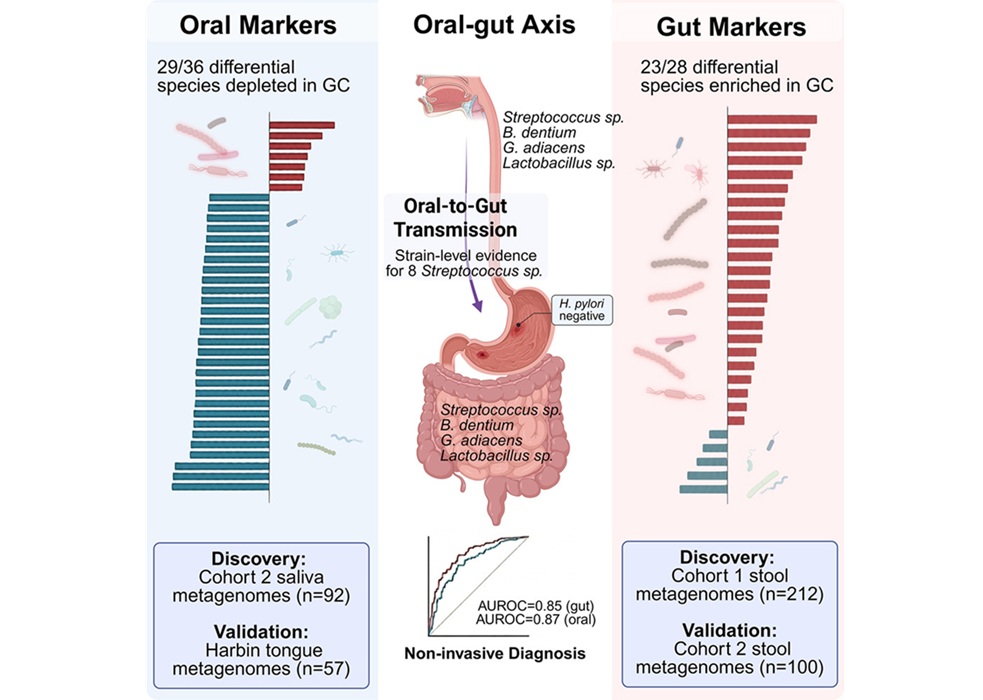
- Oral–Gut Microbiome Signatures Identify Early Gastric Cancer
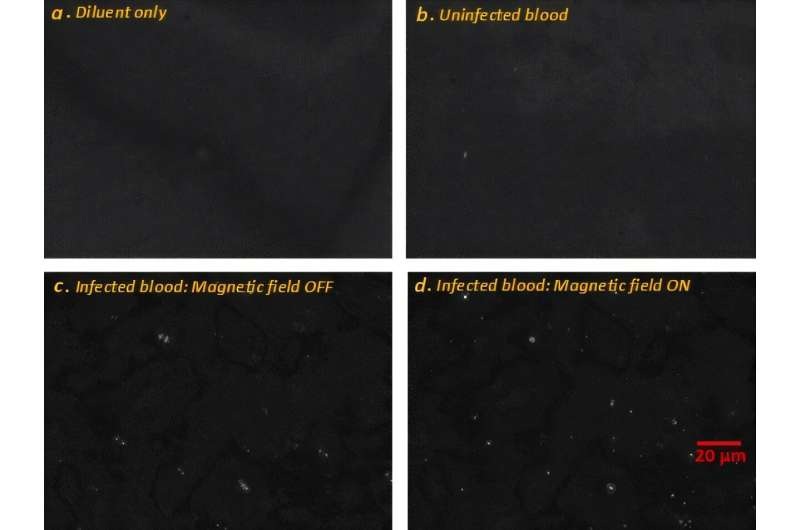
- Label-Free Microscopy Method Enables Faster, Quantitative Detection of Malaria
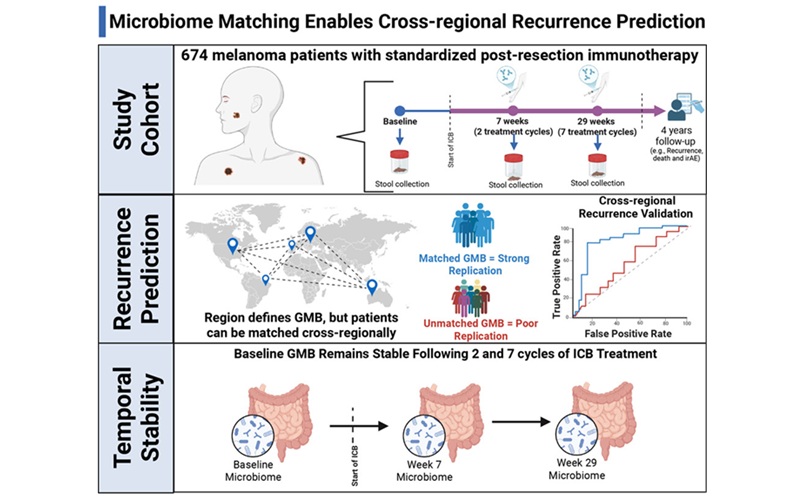
- Gut Microbiome Test Predicts Melanoma Recurrence After Surgery
- Rapid Blood-Culture Susceptibility Panel Expands Coverage for Gram-Negative Infections
- Antibiotic Resistance Genes Found in Newborns Within Hours of Birth
- New Platform Captures Extracellular Vesicles for Early Cancer Detection
- Microfluidic Single-Cell Assay Predicts Breast Cancer Risk
- AI Tool Predicts Non-Response to Targeted Therapy in Colorectal Cancer
- Integrated System Streamlines Pre-Analytical Workflow for Molecular Testing
- Noninvasive Sputum Test Detects Early Lung Cancer
- FDA Clears Blood-Based Companion Diagnostic for Advanced Breast Cancer Therapy
- CareDx Expands Precision Oncology Portfolio with Naveris Acquisition
- Thermo Fisher Scientific to Sell Microbiology Business to Astorg
- Collaboration Expands Access to Rapid Metagenomic Diagnostics for Complex Infections
- QuidelOrtho Adds Ultra-Fast PCR Platform with LEX Acquisition
- Inflammatory Gene Signature Links Metabolic Disease to Pancreatic Cancer Recurrence
- Study Links Abnormal Gene Splicing to Treatment Response in Metastatic Kidney Cancer
- Research Reveals How Some Aplastic Anemia Patients Recover Bone Marrow Function
- New Molecular Insights Support Diagnosis of Hodgkin Lymphoma
- Epigenetic Signals and Blood Markers Aid Chronic Fatigue Syndrome Diagnosis
- New AI Tool Reveals Hidden Genetic Signals in Routine H&E Slides
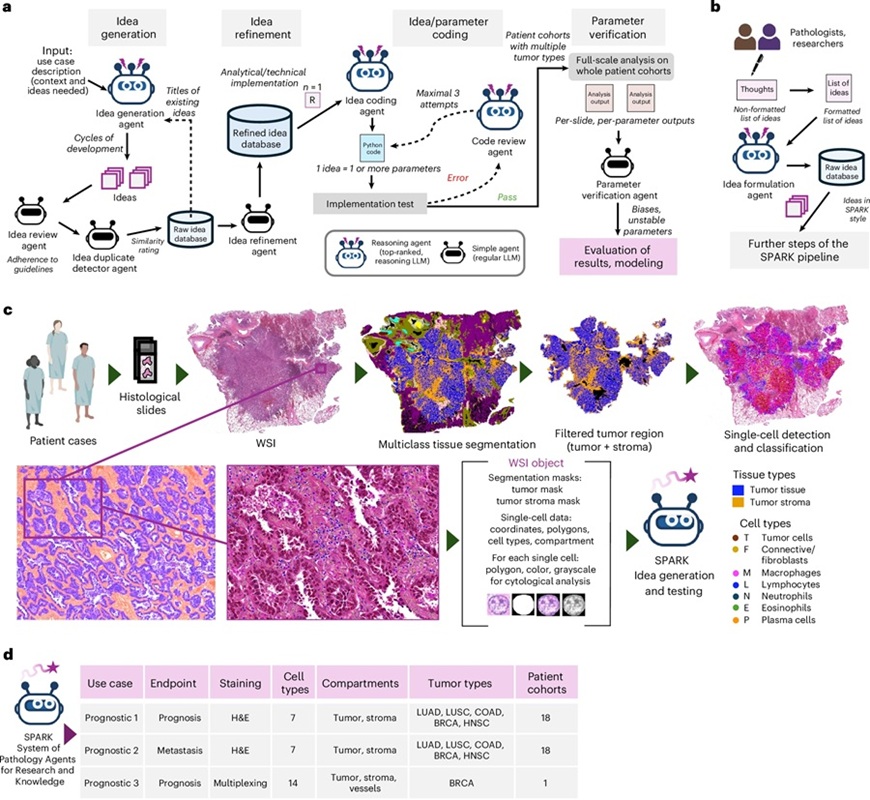
- AI System Analyzes Routine Pathology Slides to Predict Cancer Outcomes
- New Tissue Mapping Approach Identifies High-Risk Form of Diabetic Kidney Disease
- Multimodal AI Tool Predicts Genetic Alterations to Guide Breast Cancer Treatment
- Interpretable AI Reveals Hidden Cellular Features from Microscopy Images

 Expo
Expo
- Routine Blood Tests Years Before Pregnancy Could Identify Preeclampsia Risk
- Blood Test Detects Testicular Cancer Missed by Standard Markers
- Routine Blood Tests Identify Biomarkers Linked to PTSD
- Proteomic Data Underscore Need for Age-Specific Pediatric Reference Ranges
- Routine Blood Count Ratio Linked to Future Alzheimer’s and Dementia Risk
- Blood Test Maps Tumor Microenvironment to Predict Immunotherapy Response
- Multiplex Respiratory Panel Integrates Automated Extraction to Streamline High-Volume Testing
- Whole-Blood RNA Test Predicts Disease Trajectory and Treatment Response
- Blood-Based Epigenetic Test Predicts GLP-1 Response and Tracks Treatment Effects
- Tumor Genomic Testing Guides Immunotherapy Selection in Pituitary Tumors
- Blood Test Enables Early Detection of Multiple Myeloma Relapse
- Single Assay Enables Rapid HLA and ABO Genotyping for Transplant Matching
- Prognostic Biomarker Identified in Diffuse Large B-Cell Lymphoma
- Routine Blood Test Parameters Link Anemia to Cancer Risk and Mortality
- Prognostic Tool Guides Personalized Treatment in Rare Blood Cancer
- T-Cell Senescence Profiling May Predict CAR T Responses
- Finger-Prick Lateral Flow Test Detects Sepsis Biomarkers at Point of Care
- Study Highlights Low Sensitivity of Current Lyme Tests in Early Infection
- Immune Aging Clock Quantifies Immunosenescence and Identifies Therapeutic Target
- Study Finds Influenza Often Undiagnosed in Winter Deaths
- Oral–Gut Microbiome Signatures Identify Early Gastric Cancer
- Label-Free Microscopy Method Enables Faster, Quantitative Detection of Malaria
- Gut Microbiome Test Predicts Melanoma Recurrence After Surgery
- Rapid Blood-Culture Susceptibility Panel Expands Coverage for Gram-Negative Infections
- Antibiotic Resistance Genes Found in Newborns Within Hours of Birth
- New Platform Captures Extracellular Vesicles for Early Cancer Detection
- Microfluidic Single-Cell Assay Predicts Breast Cancer Risk
- AI Tool Predicts Non-Response to Targeted Therapy in Colorectal Cancer
- Integrated System Streamlines Pre-Analytical Workflow for Molecular Testing
- Noninvasive Sputum Test Detects Early Lung Cancer
- FDA Clears Blood-Based Companion Diagnostic for Advanced Breast Cancer Therapy
- CareDx Expands Precision Oncology Portfolio with Naveris Acquisition
- Thermo Fisher Scientific to Sell Microbiology Business to Astorg
- Collaboration Expands Access to Rapid Metagenomic Diagnostics for Complex Infections
- QuidelOrtho Adds Ultra-Fast PCR Platform with LEX Acquisition
- Inflammatory Gene Signature Links Metabolic Disease to Pancreatic Cancer Recurrence
- Study Links Abnormal Gene Splicing to Treatment Response in Metastatic Kidney Cancer
- Research Reveals How Some Aplastic Anemia Patients Recover Bone Marrow Function
- New Molecular Insights Support Diagnosis of Hodgkin Lymphoma
- Epigenetic Signals and Blood Markers Aid Chronic Fatigue Syndrome Diagnosis
- New AI Tool Reveals Hidden Genetic Signals in Routine H&E Slides
- AI System Analyzes Routine Pathology Slides to Predict Cancer Outcomes
- New Tissue Mapping Approach Identifies High-Risk Form of Diabetic Kidney Disease
- Multimodal AI Tool Predicts Genetic Alterations to Guide Breast Cancer Treatment
- Interpretable AI Reveals Hidden Cellular Features from Microscopy Images