Expo
view channel
view channel
view channel
view channel
view channel
view channel
view channel
view channel
view channel
Molecular DiagnosticsHematologyImmunologyMicrobiologyPathologyTechnologyIndustry
Events
Webinars

- AI Sensor Detects Neurological Disorders Using Single Saliva Drop
- New Blood Test Index Offers Earlier Detection of Liver Scarring
- Electronic Nose Smells Early Signs of Ovarian Cancer in Blood
- Simple Blood Test Offers New Path to Alzheimer’s Assessment in Primary Care
- Existing Hospital Analyzers Can Identify Fake Liquid Medical Products
- Serial Liquid Biopsies Reveal Therapy Resistance in Metastatic Prostate Cancer
- Rapid RSV Tests Reduces Antibiotic Prescribing in Kids
- DNA Aptamers Offer New Tool for Easy Alzheimer's Blood Test
- Jumping "DNA Parasites” Linked to Early Tumor Development
- AI-Based Liquid Biopsy Detects Liver Fibrosis, Cirrhosis and Chronic Disease Signals
- New Guidelines Aim to Improve AL Amyloidosis Diagnosis
- Fast and Easy Test Could Revolutionize Blood Transfusions
- Automated Hemostasis System Helps Labs of All Sizes Optimize Workflow
- High-Sensitivity Blood Test Improves Assessment of Clotting Risk in Heart Disease Patients
- AI Algorithm Effectively Distinguishes Alpha Thalassemia Subtypes
- Immune Signature Identified in Treatment-Resistant Myasthenia Gravis
- New Biomarker Predicts Chemotherapy Response in Triple-Negative Breast Cancer
- Blood Test Identifies Lung Cancer Patients Who Can Benefit from Immunotherapy Drug
- Whole-Genome Sequencing Approach Identifies Cancer Patients Benefitting From PARP-Inhibitor Treatment
- Ultrasensitive Liquid Biopsy Demonstrates Efficacy in Predicting Immunotherapy Response
- WHO Recommends Near POC Tests, Tongue Swabs and Sputum Pooling for TB Diagnosis
- New Imaging Approach Could Help Predict Dangerous Gut Infection
- Rapid Sequencing Could Transform Tuberculosis Care
- Blood-Based Viral Signature Identified in Crohn’s Disease
- Hidden Gut Viruses Linked to Colorectal Cancer Risk
- AI-Driven Diagnostic Demonstrates High Accuracy in Detecting Periprosthetic Joint Infection
- Blood Test “Clocks” Predict Start of Alzheimer’s Symptoms
- AI-Powered Biomarker Predicts Liver Cancer Risk
- Robotic Technology Unveiled for Automated Diagnostic Blood Draws
- ADLM Launches First-of-Its-Kind Data Science Program for Laboratory Medicine Professionals
- Agilent Technologies Acquires Pathology Diagnostics Company Biocare Medical
- Cepheid Joins CDC Initiative to Strengthen U.S. Pandemic Testing Preparednesss
- QuidelOrtho Collaborates with Lifotronic to Expand Global Immunoassay Portfolio
- WHX Labs in Dubai spotlights leadership skills shaping next-generation laboratories
- New Collaboration Brings Automated Mass Spectrometry to Routine Laboratory Testing
- Barcoded DNA Sheds Light on Hidden Complexities in Breast Cancer Detection
- CRISPR-Based Platform Pinpoints Drivers of Acute Myeloid Leukemia in Patient Cells
- Protective Brain Protein Emerges as Biomarker Target in Alzheimer’s Disease
- Genome Analysis Predicts Likelihood of Neurodisability in Oxygen-Deprived Newborns
- Gene Panel Predicts Disease Progession for Patients with B-cell Lymphoma
- Sex Differences in Alzheimer’s Biomarkers Linked to Faster Cognitive Decline
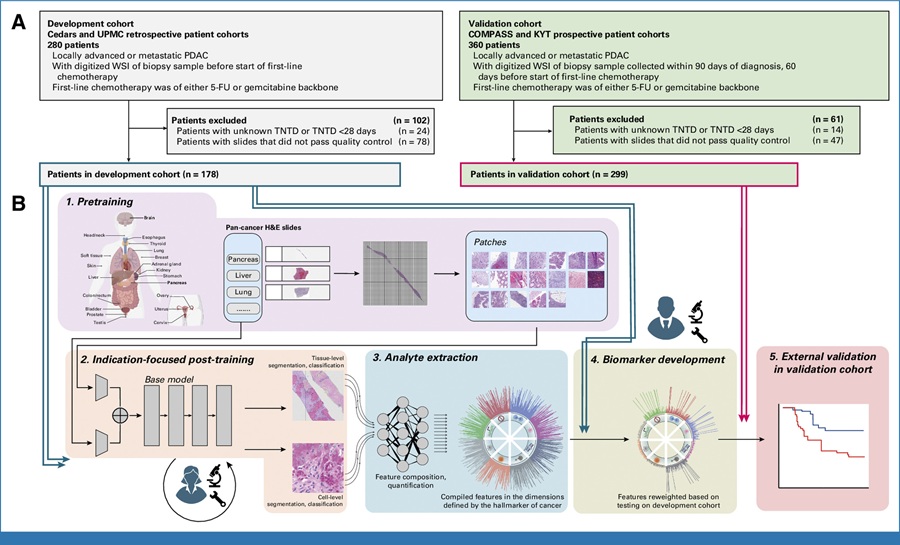
- AI Tool Predicts Chemotherapy Response from Biopsy Slides
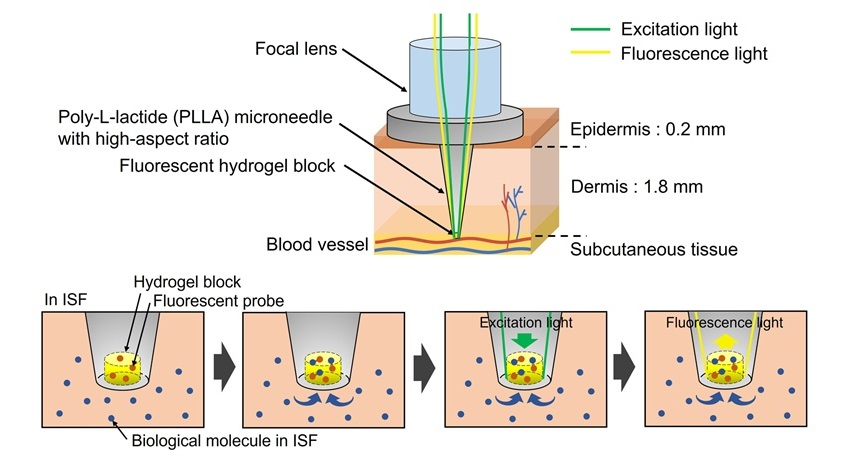
- World’s First Optical Microneedle Device to Enable Blood-Sampling-Free Clinical Testing
- Pathogen-Agnostic Testing Reveals Hidden Respiratory Threats in Negative Samples
- Molecular Imaging to Reduce Need for Melanoma Biopsies

 Expo
Expo
- AI Sensor Detects Neurological Disorders Using Single Saliva Drop
- New Blood Test Index Offers Earlier Detection of Liver Scarring
- Electronic Nose Smells Early Signs of Ovarian Cancer in Blood
- Simple Blood Test Offers New Path to Alzheimer’s Assessment in Primary Care
- Existing Hospital Analyzers Can Identify Fake Liquid Medical Products
- Serial Liquid Biopsies Reveal Therapy Resistance in Metastatic Prostate Cancer
- Rapid RSV Tests Reduces Antibiotic Prescribing in Kids
- DNA Aptamers Offer New Tool for Easy Alzheimer's Blood Test
- Jumping "DNA Parasites” Linked to Early Tumor Development
- AI-Based Liquid Biopsy Detects Liver Fibrosis, Cirrhosis and Chronic Disease Signals
- New Guidelines Aim to Improve AL Amyloidosis Diagnosis
- Fast and Easy Test Could Revolutionize Blood Transfusions
- Automated Hemostasis System Helps Labs of All Sizes Optimize Workflow
- High-Sensitivity Blood Test Improves Assessment of Clotting Risk in Heart Disease Patients
- AI Algorithm Effectively Distinguishes Alpha Thalassemia Subtypes
- Immune Signature Identified in Treatment-Resistant Myasthenia Gravis
- New Biomarker Predicts Chemotherapy Response in Triple-Negative Breast Cancer
- Blood Test Identifies Lung Cancer Patients Who Can Benefit from Immunotherapy Drug
- Whole-Genome Sequencing Approach Identifies Cancer Patients Benefitting From PARP-Inhibitor Treatment
- Ultrasensitive Liquid Biopsy Demonstrates Efficacy in Predicting Immunotherapy Response
- WHO Recommends Near POC Tests, Tongue Swabs and Sputum Pooling for TB Diagnosis
- New Imaging Approach Could Help Predict Dangerous Gut Infection
- Rapid Sequencing Could Transform Tuberculosis Care
- Blood-Based Viral Signature Identified in Crohn’s Disease
- Hidden Gut Viruses Linked to Colorectal Cancer Risk
- AI-Driven Diagnostic Demonstrates High Accuracy in Detecting Periprosthetic Joint Infection
- Blood Test “Clocks” Predict Start of Alzheimer’s Symptoms
- AI-Powered Biomarker Predicts Liver Cancer Risk
- Robotic Technology Unveiled for Automated Diagnostic Blood Draws
- ADLM Launches First-of-Its-Kind Data Science Program for Laboratory Medicine Professionals
- Agilent Technologies Acquires Pathology Diagnostics Company Biocare Medical
- Cepheid Joins CDC Initiative to Strengthen U.S. Pandemic Testing Preparednesss
- QuidelOrtho Collaborates with Lifotronic to Expand Global Immunoassay Portfolio
- WHX Labs in Dubai spotlights leadership skills shaping next-generation laboratories
- New Collaboration Brings Automated Mass Spectrometry to Routine Laboratory Testing
- Barcoded DNA Sheds Light on Hidden Complexities in Breast Cancer Detection
- CRISPR-Based Platform Pinpoints Drivers of Acute Myeloid Leukemia in Patient Cells
- Protective Brain Protein Emerges as Biomarker Target in Alzheimer’s Disease
- Genome Analysis Predicts Likelihood of Neurodisability in Oxygen-Deprived Newborns
- Gene Panel Predicts Disease Progession for Patients with B-cell Lymphoma
- Sex Differences in Alzheimer’s Biomarkers Linked to Faster Cognitive Decline
- AI Tool Predicts Chemotherapy Response from Biopsy Slides
- World’s First Optical Microneedle Device to Enable Blood-Sampling-Free Clinical Testing
- Pathogen-Agnostic Testing Reveals Hidden Respiratory Threats in Negative Samples
- Molecular Imaging to Reduce Need for Melanoma Biopsies