Expo
view channel
view channel
view channel
view channel
view channel
view channel
view channel
view channel
view channel
Clinical Chem.Molecular DiagnosticsHematologyImmunologyMicrobiology
TechnologyIndustry
Events
Webinars

- Study Finds ApoB Testing More Effective Than LDL for Guiding Lipid Therapy
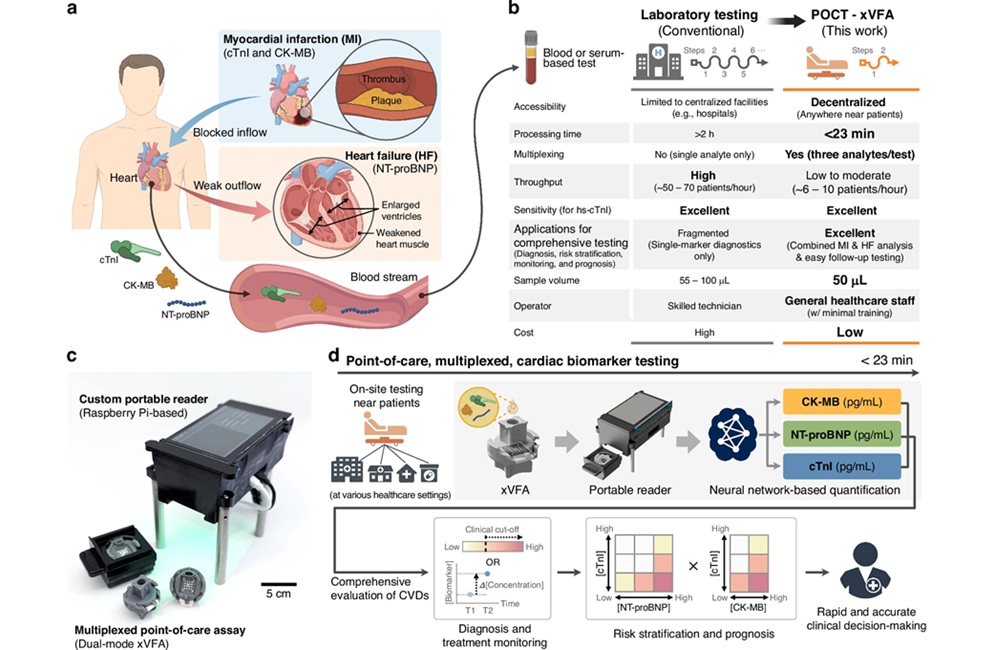
- AI-Enabled POC Test Quantifies Multiple Cardiac Biomarkers
- Next Generation Automated Analyzers Increase Throughput for Clinical Chemistry and Electrolyte Testing
- Blood Metabolite Test Detects Early Cognitive Decline
- AI-Based Blood Test Diagnose Multiple Brain Disorders from Blood Sample
- CE-Marked Blood Test Enables Monitoring of Neuroinflammation in Multiple Sclerosis
- Ultrasensitive Assay Tracks Resistance Mutations MRD Monitoring
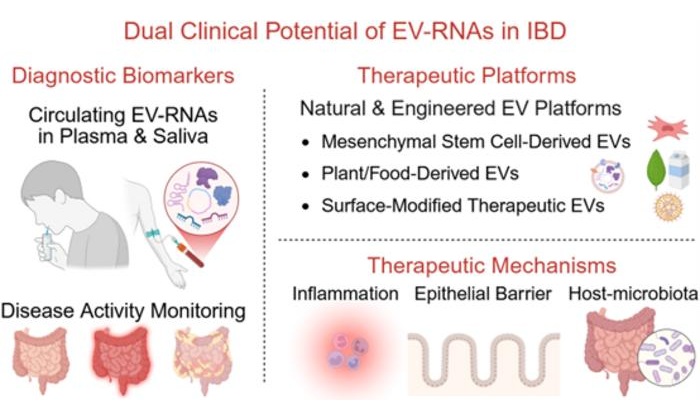
- Extracellular Vesicle RNA Biomarkers Enable Noninvasive IBD Diagnosis and Monitoring
- FDA Clears At-Home HPV Test with Extended Genotyping for Cervical Screening
- New Gene Signature Reveals Underdiagnosed Lung Cancer Subtype
- Prognostic Tool Guides Personalized Treatment in Rare Blood Cancer
- New Platelet Function Assay Enables Monitoring of Antiplatelet Therapy
- Open Multi-Omics Platform Identifies Prognostic Subtypes in Blood Cancers
- AI-Powered Digital Workflow Standardizes Bone Marrow Aspirate Morphology
- Rapid Cartridge-Based Test Aims to Expand Access to Hemoglobin Disorder Diagnosis
- Combined Screening Approach Identifies Early Leprosy Cases
- Antibody Blood Test Identifies Active TB and Distinguishes Latent Infection
- FDA Approval Expands Use of PD-L1 Companion Diagnostic in Esophageal and GEJ Carcinomas
- Study Identifies Inflammatory Pathway Driving Immunotherapy Resistance in Bladder Cancer
- Microfluidic Chip Detects Cancer Recurrence from Immune Response Signals
- Cost-Effective Sampling and Sequencing Workflow Identifies ICU Infection Hotspots
- New Bacterial Target Identified for Early Detection of Noma
- Genomic Analysis Links Emerging Streptococcal Strains to Specific Infections
- Rapid Urine Test Speeds Antibiotic Selection for UTIs
- WHO Endorses Rapid Point-of-Care Testing to Improve TB Detection
- Noninvasive Sputum Test Detects Early Lung Cancer
- New AI Tool Enables Rapid Treatment Selection in Pediatric Leukemia
- Breakthrough Mass Spectrometry Design Could Enable Ultra-Low Abundance Detection
- Rapid Biosensor Detects Drug Sensitivity in Breast Tumors
- Online Tool Supports Family Screening for Inherited Cancer Risk
- Junction, GRAIL Enable Scalable Access to Multi-Cancer Detection Test
- GRAIL Partners with Epic to Integrate Multi-Cancer Test into EHR
- Global Partnership Aims to Streamline NGS Tumor Profiling in Oncology Trials
- Takara Bio USA and Hamilton Partner to Automate NGS Library Preparation
- Lunit and CellCarta Collaborate to Expand AI Pathology in CDx Development
- Single-Cell Method Measures RNA and Proteins to Reveal Immune Responses
- Study Links Midlife Vitamin D to Lower Tau in Alzheimer's
- Common Metablolic Enzyme Could Predict Response to Cancer Immunotherapy
- Newly Identfied Genetic Variants in MND Support Prognosis and Family Testing
- International Consensus Standardizes Tumor Microbiota Detection and Reporting
- AI Improves Completeness of Complex Cancer Pathology Reports
- AI Tool Predicts Chemotherapy Response in Small Cell Lung Cancer
- Tumor-Specific Biomarker Predicts Neoadjuvant Immunotherapy Response in Gastric Cancer
- AI Tool Predicts Patient-Specific Chemotherapy Benefit in Breast Cancer
- AI-Based Pathology Model Guides Chemotherapy Decisions in Breast Cancer

 Expo
Expo
- Study Finds ApoB Testing More Effective Than LDL for Guiding Lipid Therapy
- AI-Enabled POC Test Quantifies Multiple Cardiac Biomarkers
- Next Generation Automated Analyzers Increase Throughput for Clinical Chemistry and Electrolyte Testing
- Blood Metabolite Test Detects Early Cognitive Decline
- AI-Based Blood Test Diagnose Multiple Brain Disorders from Blood Sample
- CE-Marked Blood Test Enables Monitoring of Neuroinflammation in Multiple Sclerosis
- Ultrasensitive Assay Tracks Resistance Mutations MRD Monitoring
- Extracellular Vesicle RNA Biomarkers Enable Noninvasive IBD Diagnosis and Monitoring
- FDA Clears At-Home HPV Test with Extended Genotyping for Cervical Screening
- New Gene Signature Reveals Underdiagnosed Lung Cancer Subtype
- Prognostic Tool Guides Personalized Treatment in Rare Blood Cancer
- New Platelet Function Assay Enables Monitoring of Antiplatelet Therapy
- Open Multi-Omics Platform Identifies Prognostic Subtypes in Blood Cancers
- AI-Powered Digital Workflow Standardizes Bone Marrow Aspirate Morphology
- Rapid Cartridge-Based Test Aims to Expand Access to Hemoglobin Disorder Diagnosis
- Combined Screening Approach Identifies Early Leprosy Cases
- Antibody Blood Test Identifies Active TB and Distinguishes Latent Infection
- FDA Approval Expands Use of PD-L1 Companion Diagnostic in Esophageal and GEJ Carcinomas
- Study Identifies Inflammatory Pathway Driving Immunotherapy Resistance in Bladder Cancer
- Microfluidic Chip Detects Cancer Recurrence from Immune Response Signals
- Cost-Effective Sampling and Sequencing Workflow Identifies ICU Infection Hotspots
- New Bacterial Target Identified for Early Detection of Noma
- Genomic Analysis Links Emerging Streptococcal Strains to Specific Infections
- Rapid Urine Test Speeds Antibiotic Selection for UTIs
- WHO Endorses Rapid Point-of-Care Testing to Improve TB Detection
- Noninvasive Sputum Test Detects Early Lung Cancer
- New AI Tool Enables Rapid Treatment Selection in Pediatric Leukemia
- Breakthrough Mass Spectrometry Design Could Enable Ultra-Low Abundance Detection
- Rapid Biosensor Detects Drug Sensitivity in Breast Tumors
- Online Tool Supports Family Screening for Inherited Cancer Risk
- Junction, GRAIL Enable Scalable Access to Multi-Cancer Detection Test
- GRAIL Partners with Epic to Integrate Multi-Cancer Test into EHR
- Global Partnership Aims to Streamline NGS Tumor Profiling in Oncology Trials
- Takara Bio USA and Hamilton Partner to Automate NGS Library Preparation
- Lunit and CellCarta Collaborate to Expand AI Pathology in CDx Development
- Single-Cell Method Measures RNA and Proteins to Reveal Immune Responses
- Study Links Midlife Vitamin D to Lower Tau in Alzheimer's
- Common Metablolic Enzyme Could Predict Response to Cancer Immunotherapy
- Newly Identfied Genetic Variants in MND Support Prognosis and Family Testing
- International Consensus Standardizes Tumor Microbiota Detection and Reporting
- AI Improves Completeness of Complex Cancer Pathology Reports
- AI Tool Predicts Chemotherapy Response in Small Cell Lung Cancer
- Tumor-Specific Biomarker Predicts Neoadjuvant Immunotherapy Response in Gastric Cancer
- AI Tool Predicts Patient-Specific Chemotherapy Benefit in Breast Cancer
- AI-Based Pathology Model Guides Chemotherapy Decisions in Breast Cancer