Expo
view channel
view channel
view channel
view channel
view channel
view channel
view channel
view channel
view channel
Molecular DiagnosticsHematologyImmunologyMicrobiologyPathologyTechnologyIndustry
Events
Webinars

- Automated NfL Assay Supports Monitoring of Neurological Disorders
- CSF Biomarker Improves Diagnosis of Parkinson’s Disease and Lewy Body Dementia
- Simple Urine Home Test Kit Could Detect Early-Stage Breast Cancer
- New Tool Tracks Biomarker Changes to Predict Myeloma Progression
- New Plasma Tau Assay Improves Prediction of Alzheimer’s Progression
- Simple Nasal Swab May Reveal Early Signs of Alzheimer’s Disease
- Blood Biomarker Predicts Cognitive Outcomes After Cardiac Arrest
- Liquid Biopsy Enables Faster Diagnosis of Childhood Cancer in Africa
- Blood Test Helps Guide Treatment in Older Women with Breast Cancer
- Liquid Biopsy Method Pinpoints Disease Source From a Single Drop of Blood
- New Guidelines Aim to Improve AL Amyloidosis Diagnosis
- Fast and Easy Test Could Revolutionize Blood Transfusions
- Automated Hemostasis System Helps Labs of All Sizes Optimize Workflow
- High-Sensitivity Blood Test Improves Assessment of Clotting Risk in Heart Disease Patients
- AI Algorithm Effectively Distinguishes Alpha Thalassemia Subtypes
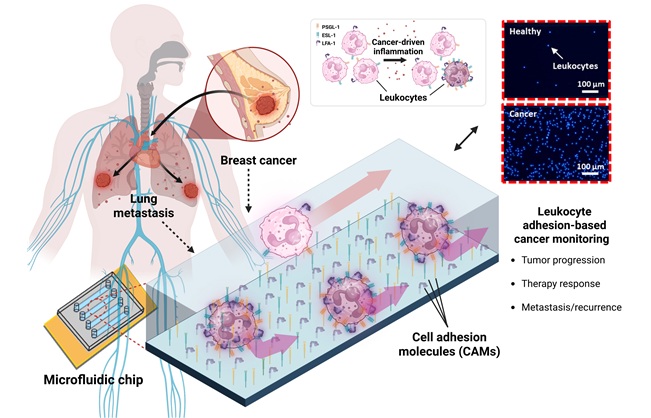
- Microfluidic Chip Detects Cancer Recurrence from Immune Response Signals
- Cancer Mutation ‘Fingerprints’ to Improve Prediction of Immunotherapy Response
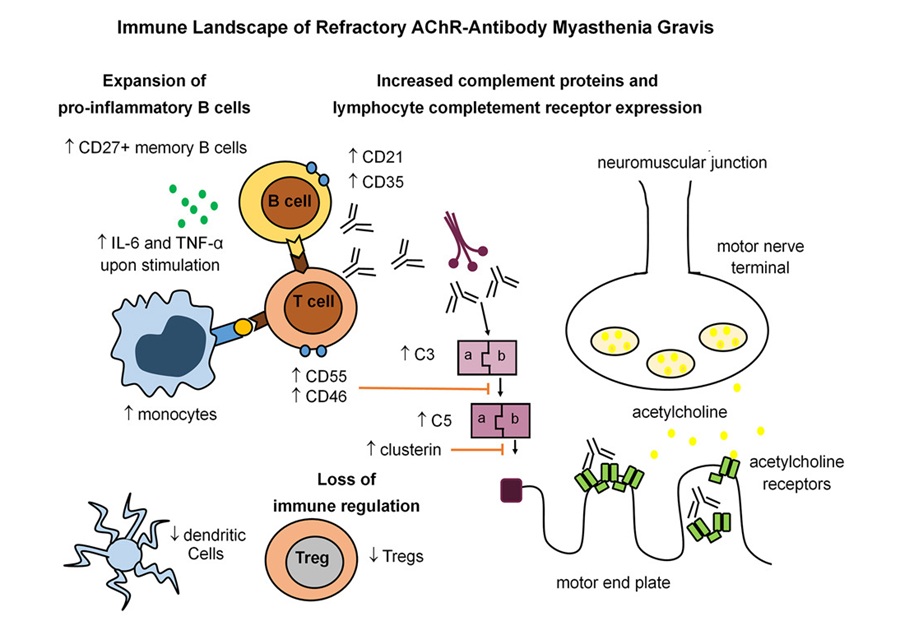
- Immune Signature Identified in Treatment-Resistant Myasthenia Gravis
- New Biomarker Predicts Chemotherapy Response in Triple-Negative Breast Cancer
- Blood Test Identifies Lung Cancer Patients Who Can Benefit from Immunotherapy Drug
- Study Highlights Accuracy Gaps in Consumer Gut Microbiome Kits
- WHO Recommends Near POC Tests, Tongue Swabs and Sputum Pooling for TB Diagnosis
- New Imaging Approach Could Help Predict Dangerous Gut Infection
- Rapid Sequencing Could Transform Tuberculosis Care
- Blood-Based Viral Signature Identified in Crohn’s Disease
- Portable Breath Sensor Detects Pneumonia Biomarkers in Minutes
- New Electronic Pipette Enhances Workflows with Touchscreen Control
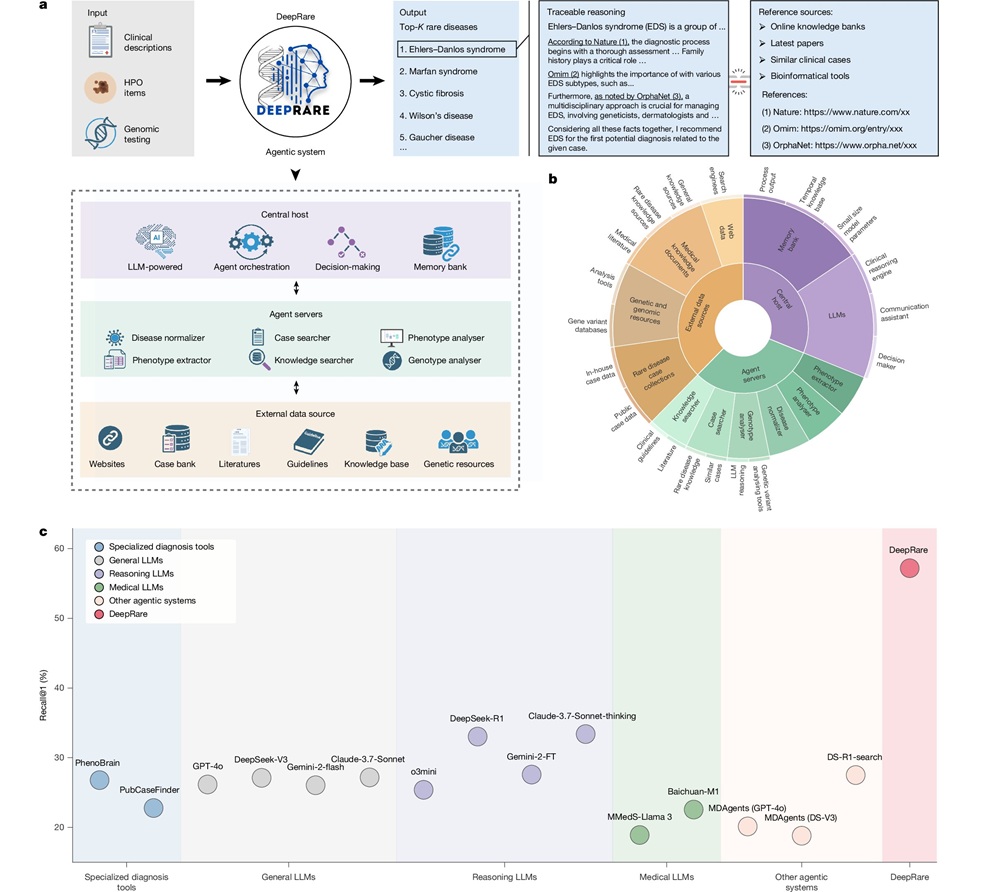
- AI Model Outperforms Clinicians in Rare Disease Detection
- AI-Driven Diagnostic Demonstrates High Accuracy in Detecting Periprosthetic Joint Infection
- Blood Test “Clocks” Predict Start of Alzheimer’s Symptoms
- Co-Diagnostics Agreement Expands Commercial and Distribution Reach in South Asia
- Automated MSI Test Gains IVDR Certification to Guide CRC Therapy
- New Partnership Brings Alzheimer’s Blood Biomarker Test to Community Screening Network
- MGI Tech Strengthens Sequencing Portfolio with Dual Acquisition
- Agilent Technologies Acquires Pathology Diagnostics Company Biocare Medical
- Genetic Mutation Behind Aggressive Adult Leukemia Offers Treatment Clues
- Disease Gene Discovery Advances Diagnosis of Rare Movement Disorders
- Genetic Discovery Could Improve Diagnosis of Drug-Resistant Epilepsy
- Genetic Discovery May Improve Diagnosis of Rare Dementia Subtype
- Mass Spectrometry Technique Detects Protein and Sugar Changes in Neurodegeneration
- AI-Powered Tool to Transform Dermatopathology Workflow
- New Chromogenic Culture Media Enable Rapid Detection of Candida Infections
- Novel mcPCR Technology to Transform Testing of Clinical Samples
- Sex Differences in Alzheimer’s Biomarkers Linked to Faster Cognitive Decline
- AI Tool Predicts Chemotherapy Response from Biopsy Slides

 Expo
Expo
- Automated NfL Assay Supports Monitoring of Neurological Disorders
- CSF Biomarker Improves Diagnosis of Parkinson’s Disease and Lewy Body Dementia
- Simple Urine Home Test Kit Could Detect Early-Stage Breast Cancer
- New Tool Tracks Biomarker Changes to Predict Myeloma Progression
- New Plasma Tau Assay Improves Prediction of Alzheimer’s Progression
- Simple Nasal Swab May Reveal Early Signs of Alzheimer’s Disease
- Blood Biomarker Predicts Cognitive Outcomes After Cardiac Arrest
- Liquid Biopsy Enables Faster Diagnosis of Childhood Cancer in Africa
- Blood Test Helps Guide Treatment in Older Women with Breast Cancer
- Liquid Biopsy Method Pinpoints Disease Source From a Single Drop of Blood
- New Guidelines Aim to Improve AL Amyloidosis Diagnosis
- Fast and Easy Test Could Revolutionize Blood Transfusions
- Automated Hemostasis System Helps Labs of All Sizes Optimize Workflow
- High-Sensitivity Blood Test Improves Assessment of Clotting Risk in Heart Disease Patients
- AI Algorithm Effectively Distinguishes Alpha Thalassemia Subtypes
- Microfluidic Chip Detects Cancer Recurrence from Immune Response Signals
- Cancer Mutation ‘Fingerprints’ to Improve Prediction of Immunotherapy Response
- Immune Signature Identified in Treatment-Resistant Myasthenia Gravis
- New Biomarker Predicts Chemotherapy Response in Triple-Negative Breast Cancer
- Blood Test Identifies Lung Cancer Patients Who Can Benefit from Immunotherapy Drug
- Study Highlights Accuracy Gaps in Consumer Gut Microbiome Kits
- WHO Recommends Near POC Tests, Tongue Swabs and Sputum Pooling for TB Diagnosis
- New Imaging Approach Could Help Predict Dangerous Gut Infection
- Rapid Sequencing Could Transform Tuberculosis Care
- Blood-Based Viral Signature Identified in Crohn’s Disease
- Portable Breath Sensor Detects Pneumonia Biomarkers in Minutes
- New Electronic Pipette Enhances Workflows with Touchscreen Control
- AI Model Outperforms Clinicians in Rare Disease Detection
- AI-Driven Diagnostic Demonstrates High Accuracy in Detecting Periprosthetic Joint Infection
- Blood Test “Clocks” Predict Start of Alzheimer’s Symptoms
- Co-Diagnostics Agreement Expands Commercial and Distribution Reach in South Asia
- Automated MSI Test Gains IVDR Certification to Guide CRC Therapy
- New Partnership Brings Alzheimer’s Blood Biomarker Test to Community Screening Network
- MGI Tech Strengthens Sequencing Portfolio with Dual Acquisition
- Agilent Technologies Acquires Pathology Diagnostics Company Biocare Medical
- Genetic Mutation Behind Aggressive Adult Leukemia Offers Treatment Clues
- Disease Gene Discovery Advances Diagnosis of Rare Movement Disorders
- Genetic Discovery Could Improve Diagnosis of Drug-Resistant Epilepsy
- Genetic Discovery May Improve Diagnosis of Rare Dementia Subtype
- Mass Spectrometry Technique Detects Protein and Sugar Changes in Neurodegeneration
- AI-Powered Tool to Transform Dermatopathology Workflow
- New Chromogenic Culture Media Enable Rapid Detection of Candida Infections
- Novel mcPCR Technology to Transform Testing of Clinical Samples
- Sex Differences in Alzheimer’s Biomarkers Linked to Faster Cognitive Decline
- AI Tool Predicts Chemotherapy Response from Biopsy Slides