Expo
view channel
view channel
view channel
view channel
view channel
view channel
view channel
view channel
view channel
Clinical Chem.Molecular DiagnosticsHematology
MicrobiologyPathologyTechnologyIndustry
Events
Webinars

- Routine Blood Tests Identify Biomarkers Linked to PTSD
- Proteomic Data Underscore Need for Age-Specific Pediatric Reference Ranges
- Routine Blood Count Ratio Linked to Future Alzheimer’s and Dementia Risk
- Label-Free Microfluidic Device Enriches Tumor Cells and Clusters from Pleural Effusions
- Rapid Biosensor Detects Pancreatic Cancer Biomarker for Early Detection
- Multi-Omic Assay Predicts Recurrence and Radiation Benefit in Early Breast Cancer
- Portable Test Detects Tuberculosis from Tongue Swabs in 30 Minutes
- Blood Test Receives FDA Breakthrough Status to Differentiate Schizophrenia and Bipolar Disorder
- Genomic Risk Score Identifies Inherited Risk for Multiple Cardiovascular Conditions
- Routine Genetic Marker May Help Guide Targeted Therapy in Acute Leukemia
- Blood Test Enables Early Detection of Multiple Myeloma Relapse
- Single Assay Enables Rapid HLA and ABO Genotyping for Transplant Matching
- Prognostic Biomarker Identified in Diffuse Large B-Cell Lymphoma
- Routine Blood Test Parameters Link Anemia to Cancer Risk and Mortality
- Prognostic Tool Guides Personalized Treatment in Rare Blood Cancer
- Study Highlights Low Sensitivity of Current Lyme Tests in Early Infection
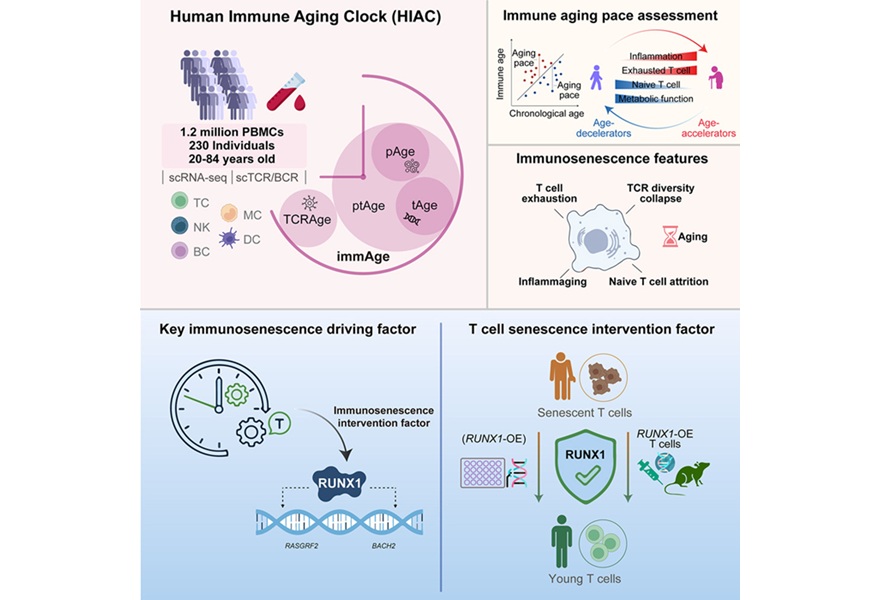
- Immune Aging Clock Quantifies Immunosenescence and Identifies Therapeutic Target
- Study Finds Influenza Often Undiagnosed in Winter Deaths
- Combined Screening Approach Identifies Early Leprosy Cases
- Antibody Blood Test Identifies Active TB and Distinguishes Latent Infection
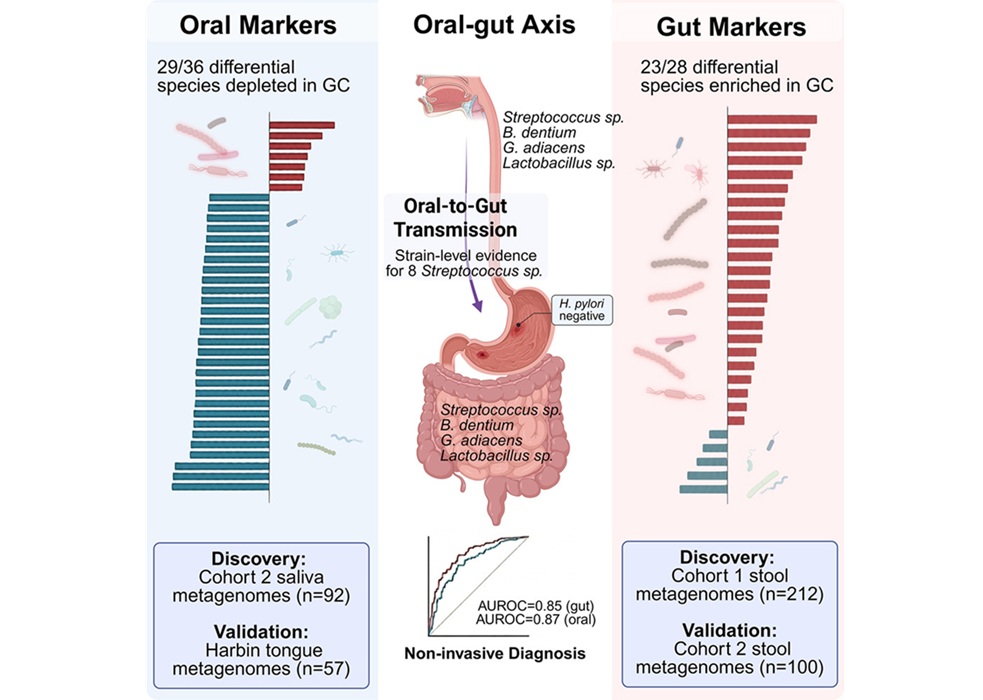
- Oral–Gut Microbiome Signatures Identify Early Gastric Cancer
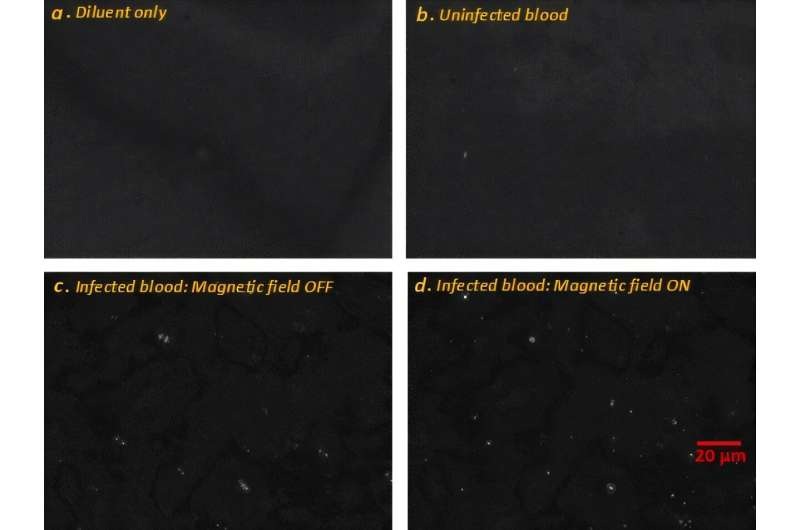
- Label-Free Microscopy Method Enables Faster, Quantitative Detection of Malaria
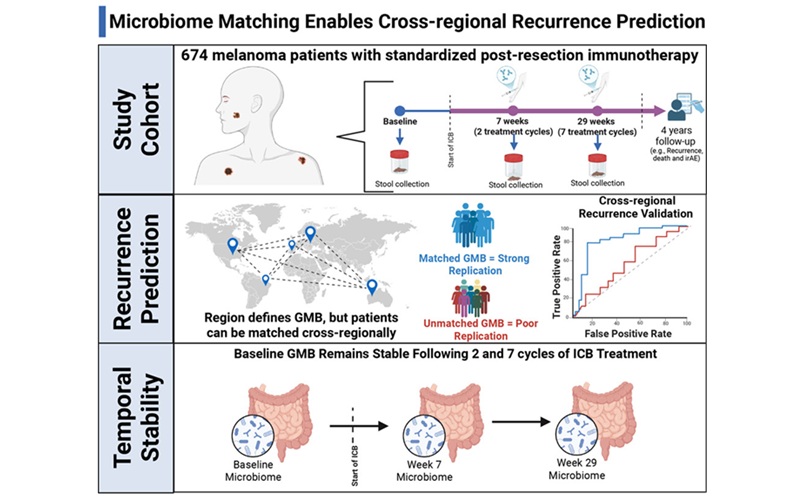
- Gut Microbiome Test Predicts Melanoma Recurrence After Surgery
- Rapid Blood-Culture Susceptibility Panel Expands Coverage for Gram-Negative Infections
- Antibiotic Resistance Genes Found in Newborns Within Hours of Birth
- AI Tool Predicts Non-Response to Targeted Therapy in Colorectal Cancer
- Integrated System Streamlines Pre-Analytical Workflow for Molecular Testing
- Noninvasive Sputum Test Detects Early Lung Cancer
- New AI Tool Enables Rapid Treatment Selection in Pediatric Leukemia
- Breakthrough Mass Spectrometry Design Could Enable Ultra-Low Abundance Detection
- Thermo Fisher Scientific to Sell Microbiology Business to Astorg
- Collaboration Expands Access to Rapid Metagenomic Diagnostics for Complex Infections
- QuidelOrtho Adds Ultra-Fast PCR Platform with LEX Acquisition
- Seegene Showcases Real-Time PCR Data Analytics Platform at ESCMID
- Roche Affiliate Expands MRD Portfolio with SAGA Acquisition
- Epigenetic Signals and Blood Markers Aid Chronic Fatigue Syndrome Diagnosis
- Microenvironment Biomarkers Could Enable Early Lung Cancer Detection
- Study Identifies Protein Changes Driving Immunotherapy Resistance in Multiple Myeloma
- Genetic Analysis Identifies BRCA-Linked Risks Across Multiple Cancers
- Study Identifies Hidden B-Cell Mutations in Autoimmune Disease
- Multimodal AI Tool Predicts Genetic Alterations to Guide Breast Cancer Treatment
- Interpretable AI Reveals Hidden Cellular Features from Microscopy Images
- Tumor Immune Structure Predicts Response to Immunotherapy in Melanoma
- Plug-and-Play AI Pathology System Classifies Multiple Cancers from Few Slides
- AI-Based Assays Support Risk Stratification in Prostate and Breast Cancer

 Expo
Expo
- Routine Blood Tests Identify Biomarkers Linked to PTSD
- Proteomic Data Underscore Need for Age-Specific Pediatric Reference Ranges
- Routine Blood Count Ratio Linked to Future Alzheimer’s and Dementia Risk
- Label-Free Microfluidic Device Enriches Tumor Cells and Clusters from Pleural Effusions
- Rapid Biosensor Detects Pancreatic Cancer Biomarker for Early Detection
- Multi-Omic Assay Predicts Recurrence and Radiation Benefit in Early Breast Cancer
- Portable Test Detects Tuberculosis from Tongue Swabs in 30 Minutes
- Blood Test Receives FDA Breakthrough Status to Differentiate Schizophrenia and Bipolar Disorder
- Genomic Risk Score Identifies Inherited Risk for Multiple Cardiovascular Conditions
- Routine Genetic Marker May Help Guide Targeted Therapy in Acute Leukemia
- Blood Test Enables Early Detection of Multiple Myeloma Relapse
- Single Assay Enables Rapid HLA and ABO Genotyping for Transplant Matching
- Prognostic Biomarker Identified in Diffuse Large B-Cell Lymphoma
- Routine Blood Test Parameters Link Anemia to Cancer Risk and Mortality
- Prognostic Tool Guides Personalized Treatment in Rare Blood Cancer
- Study Highlights Low Sensitivity of Current Lyme Tests in Early Infection
- Immune Aging Clock Quantifies Immunosenescence and Identifies Therapeutic Target
- Study Finds Influenza Often Undiagnosed in Winter Deaths
- Combined Screening Approach Identifies Early Leprosy Cases
- Antibody Blood Test Identifies Active TB and Distinguishes Latent Infection
- Oral–Gut Microbiome Signatures Identify Early Gastric Cancer
- Label-Free Microscopy Method Enables Faster, Quantitative Detection of Malaria
- Gut Microbiome Test Predicts Melanoma Recurrence After Surgery
- Rapid Blood-Culture Susceptibility Panel Expands Coverage for Gram-Negative Infections
- Antibiotic Resistance Genes Found in Newborns Within Hours of Birth
- AI Tool Predicts Non-Response to Targeted Therapy in Colorectal Cancer
- Integrated System Streamlines Pre-Analytical Workflow for Molecular Testing
- Noninvasive Sputum Test Detects Early Lung Cancer
- New AI Tool Enables Rapid Treatment Selection in Pediatric Leukemia
- Breakthrough Mass Spectrometry Design Could Enable Ultra-Low Abundance Detection
- Thermo Fisher Scientific to Sell Microbiology Business to Astorg
- Collaboration Expands Access to Rapid Metagenomic Diagnostics for Complex Infections
- QuidelOrtho Adds Ultra-Fast PCR Platform with LEX Acquisition
- Seegene Showcases Real-Time PCR Data Analytics Platform at ESCMID
- Roche Affiliate Expands MRD Portfolio with SAGA Acquisition
- Epigenetic Signals and Blood Markers Aid Chronic Fatigue Syndrome Diagnosis
- Microenvironment Biomarkers Could Enable Early Lung Cancer Detection
- Study Identifies Protein Changes Driving Immunotherapy Resistance in Multiple Myeloma
- Genetic Analysis Identifies BRCA-Linked Risks Across Multiple Cancers
- Study Identifies Hidden B-Cell Mutations in Autoimmune Disease
- Multimodal AI Tool Predicts Genetic Alterations to Guide Breast Cancer Treatment
- Interpretable AI Reveals Hidden Cellular Features from Microscopy Images
- Tumor Immune Structure Predicts Response to Immunotherapy in Melanoma
- Plug-and-Play AI Pathology System Classifies Multiple Cancers from Few Slides
- AI-Based Assays Support Risk Stratification in Prostate and Breast Cancer