Expo
view channel
view channel
view channel
view channel
view channel
view channel
view channel
view channel
view channel
Clinical Chem.
HematologyImmunologyMicrobiologyPathologyTechnologyIndustry
Events
Webinars

- Blood-Based Alzheimer’s Test Gains CE Mark for Amyloid Pathology Detection
- FDA-Cleared Assay Enables Comprehensive Automated Testosterone Testing
- CE-Marked Blood Biomarker Test Advances Automated Alzheimer’s Diagnostics
- Noninvasive Urine Test May Support Earlier Diagnosis of Psychiatric Disorders
- At-Home Blood and Cognitive Tests Support Dementia Risk Stratification
- FDA-Approved MRD Blood Test Guides Adjuvant Bladder Cancer Therapy
- Tumor Genomic Profiling Identifies High-Risk Gallbladder Cancer
- Genome Sequencing Uncovers Hidden Genetic Risks in Healthy Adults
- Sensitive Protein Marker Aids Diagnosis of Small Cell Prostate Cancer
- Rapid Multiplex PCR Test Detects 11 Gastrointestinal Pathogens from Single Sample
- Advanced CBC-Derived Indices Integrated into Hematology Platforms
- Blood Test Enables Early Detection of Multiple Myeloma Relapse
- Single Assay Enables Rapid HLA and ABO Genotyping for Transplant Matching
- Prognostic Biomarker Identified in Diffuse Large B-Cell Lymphoma
- Routine Blood Test Parameters Link Anemia to Cancer Risk and Mortality
- Biomarkers and Molecular Testing Advance Precision Allergy Care
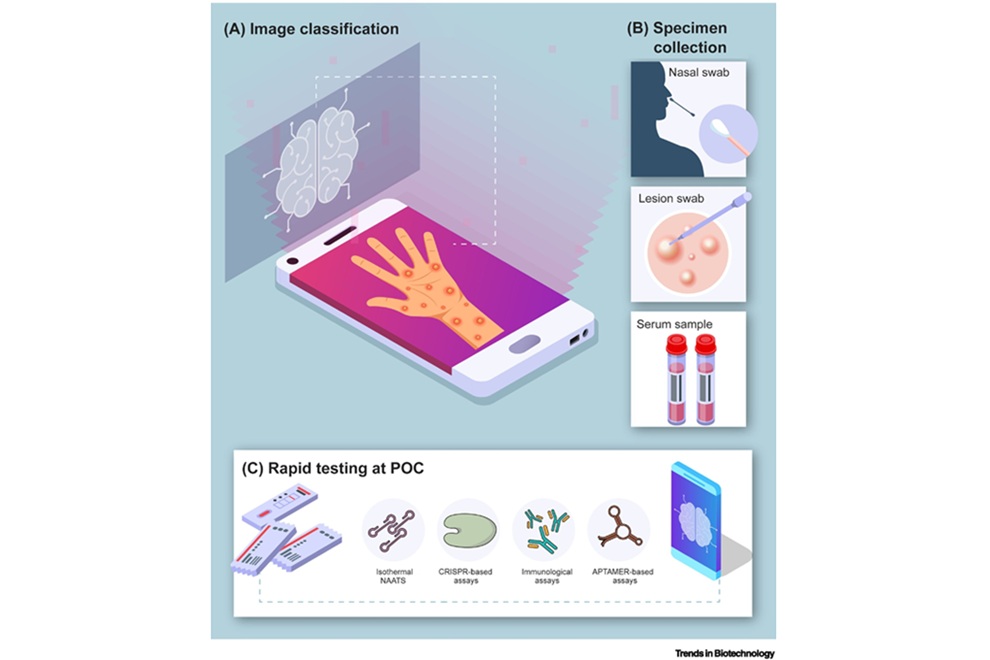
- Point-of-Care Tests Could Expand Access to Mpox Diagnosis
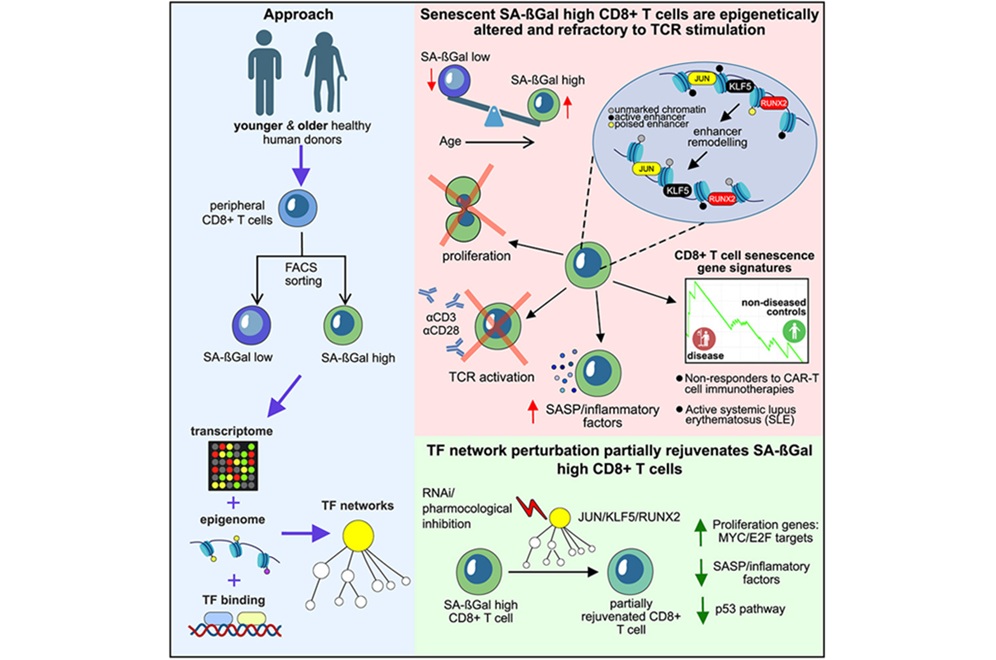
- T-Cell Senescence Profiling May Predict CAR T Responses
- Finger-Prick Lateral Flow Test Detects Sepsis Biomarkers at Point of Care
- Study Highlights Low Sensitivity of Current Lyme Tests in Early Infection
- Large-Scale Genomic Surveillance Tracks Resistant Bacteria Across European Hospitals
- Molecular Urine and Stool Tests Do Not Improve Early TB Treatment in Hospitalized HIV Patients
- Rapid Antigen Biosensor Detects Active Tuberculosis in One Hour
- Oral–Gut Microbiome Signatures Identify Early Gastric Cancer
- Label-Free Microscopy Method Enables Faster, Quantitative Detection of Malaria
- Fully Automated Sample-to-Insight Workflow Advances Latent TB Testing
- Tumor-on-a-Chip Platform Models Pancreatic Cancer Treatment Response
- New Platform Captures Extracellular Vesicles for Early Cancer Detection
- Microfluidic Single-Cell Assay Predicts Breast Cancer Risk
- AI Tool Predicts Non-Response to Targeted Therapy in Colorectal Cancer
- Partnership Aims to Bring Risk-Guided CKD Care to Health Systems
- Roche to Acquire PathAI for Up to $1.05 Billion to Strengthen AI Diagnostics Portfolio
- FDA Clears Blood-Based Companion Diagnostic for Advanced Breast Cancer Therapy
- CareDx Expands Precision Oncology Portfolio with Naveris Acquisition
- Thermo Fisher Scientific to Sell Microbiology Business to Astorg
- Genetic Markers May Help Predict Amputation Risk in Peripheral Artery Disease
- Hidden Immune Gene Defect May Explain Kaposi Sarcoma Susceptibility
- Gene Signature Shows Promise for Depression Biomarker Testing
- AI-Driven Tumor Profiling Initiative Targets Precision Therapy Development
- Researchers Map Protein and Glycosylation Across 15 Human Body Fluids
- AI Pathology Test Receives FDA Breakthrough for Bladder Cancer Risk Stratification
- FDA Clears AI Digital Pathology Tool for Breast Cancer Risk Stratification
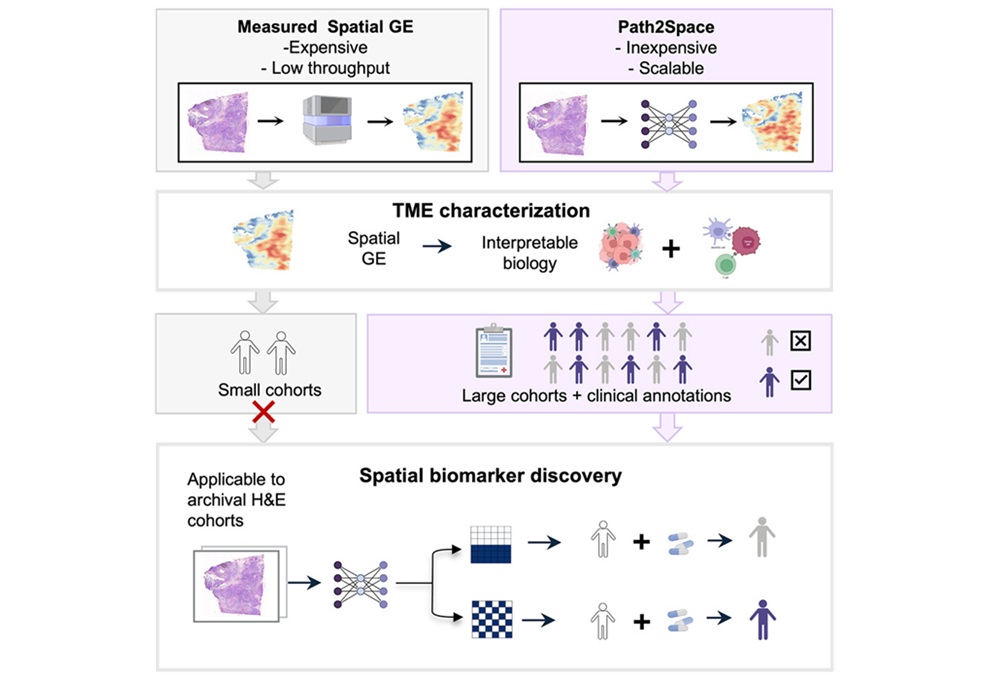
- New AI Tool Reveals Hidden Genetic Signals in Routine H&E Slides
- AI System Analyzes Routine Pathology Slides to Predict Cancer Outcomes
- New Tissue Mapping Approach Identifies High-Risk Form of Diabetic Kidney Disease

 Expo
Expo
- Blood-Based Alzheimer’s Test Gains CE Mark for Amyloid Pathology Detection
- FDA-Cleared Assay Enables Comprehensive Automated Testosterone Testing
- CE-Marked Blood Biomarker Test Advances Automated Alzheimer’s Diagnostics
- Noninvasive Urine Test May Support Earlier Diagnosis of Psychiatric Disorders
- At-Home Blood and Cognitive Tests Support Dementia Risk Stratification
- FDA-Approved MRD Blood Test Guides Adjuvant Bladder Cancer Therapy
- Tumor Genomic Profiling Identifies High-Risk Gallbladder Cancer
- Genome Sequencing Uncovers Hidden Genetic Risks in Healthy Adults
- Sensitive Protein Marker Aids Diagnosis of Small Cell Prostate Cancer
- Rapid Multiplex PCR Test Detects 11 Gastrointestinal Pathogens from Single Sample
- Advanced CBC-Derived Indices Integrated into Hematology Platforms
- Blood Test Enables Early Detection of Multiple Myeloma Relapse
- Single Assay Enables Rapid HLA and ABO Genotyping for Transplant Matching
- Prognostic Biomarker Identified in Diffuse Large B-Cell Lymphoma
- Routine Blood Test Parameters Link Anemia to Cancer Risk and Mortality
- Biomarkers and Molecular Testing Advance Precision Allergy Care
- Point-of-Care Tests Could Expand Access to Mpox Diagnosis
- T-Cell Senescence Profiling May Predict CAR T Responses
- Finger-Prick Lateral Flow Test Detects Sepsis Biomarkers at Point of Care
- Study Highlights Low Sensitivity of Current Lyme Tests in Early Infection
- Large-Scale Genomic Surveillance Tracks Resistant Bacteria Across European Hospitals
- Molecular Urine and Stool Tests Do Not Improve Early TB Treatment in Hospitalized HIV Patients
- Rapid Antigen Biosensor Detects Active Tuberculosis in One Hour
- Oral–Gut Microbiome Signatures Identify Early Gastric Cancer
- Label-Free Microscopy Method Enables Faster, Quantitative Detection of Malaria
- Fully Automated Sample-to-Insight Workflow Advances Latent TB Testing
- Tumor-on-a-Chip Platform Models Pancreatic Cancer Treatment Response
- New Platform Captures Extracellular Vesicles for Early Cancer Detection
- Microfluidic Single-Cell Assay Predicts Breast Cancer Risk
- AI Tool Predicts Non-Response to Targeted Therapy in Colorectal Cancer
- Partnership Aims to Bring Risk-Guided CKD Care to Health Systems
- Roche to Acquire PathAI for Up to $1.05 Billion to Strengthen AI Diagnostics Portfolio
- FDA Clears Blood-Based Companion Diagnostic for Advanced Breast Cancer Therapy
- CareDx Expands Precision Oncology Portfolio with Naveris Acquisition
- Thermo Fisher Scientific to Sell Microbiology Business to Astorg
- Genetic Markers May Help Predict Amputation Risk in Peripheral Artery Disease
- Hidden Immune Gene Defect May Explain Kaposi Sarcoma Susceptibility
- Gene Signature Shows Promise for Depression Biomarker Testing
- AI-Driven Tumor Profiling Initiative Targets Precision Therapy Development
- Researchers Map Protein and Glycosylation Across 15 Human Body Fluids
- AI Pathology Test Receives FDA Breakthrough for Bladder Cancer Risk Stratification
- FDA Clears AI Digital Pathology Tool for Breast Cancer Risk Stratification
- New AI Tool Reveals Hidden Genetic Signals in Routine H&E Slides
- AI System Analyzes Routine Pathology Slides to Predict Cancer Outcomes
- New Tissue Mapping Approach Identifies High-Risk Form of Diabetic Kidney Disease