Expo
view channel
view channel
view channel
view channel
view channel
view channel
view channel
view channel
view channel
Clinical Chem.Molecular Diagnostics
ImmunologyMicrobiologyPathologyTechnologyIndustry
Events
Webinars

- Blood Test Predicts Alzheimer Disease Risk Before Imaging Changes and Symptoms
- Study Finds ApoB Testing More Effective Than LDL for Guiding Lipid Therapy
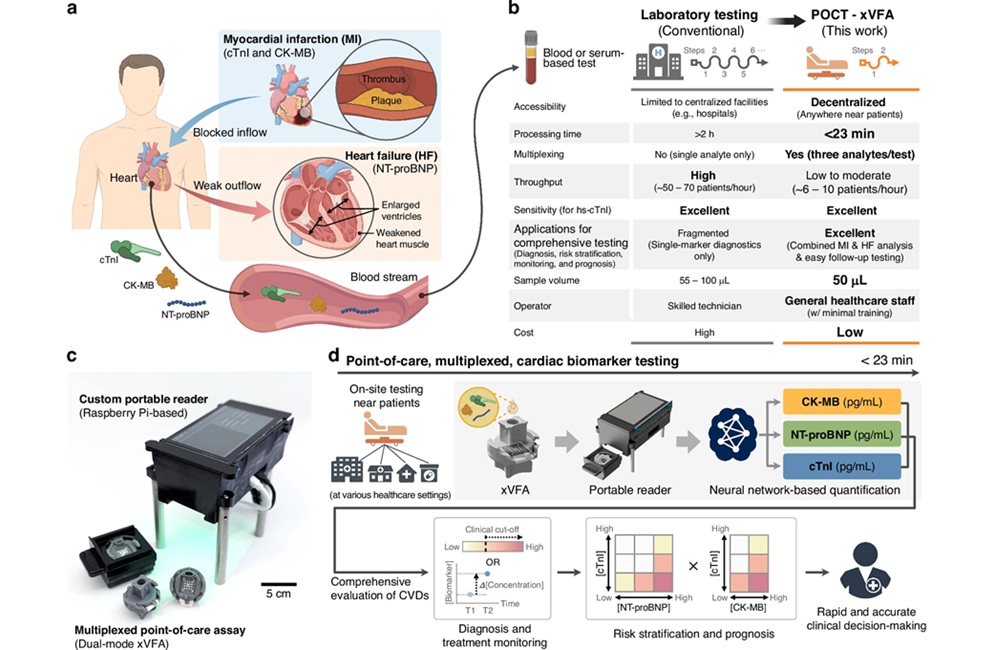
- AI-Enabled POC Test Quantifies Multiple Cardiac Biomarkers
- Next Generation Automated Analyzers Increase Throughput for Clinical Chemistry and Electrolyte Testing
- Blood Metabolite Test Detects Early Cognitive Decline
- Composite Blood Biomarkers Enable Early Detection of Common Cancers
- Machine Learning Model Uses DNA Methylation to Predict Tumor Origin in Cancers of Unknown Primary
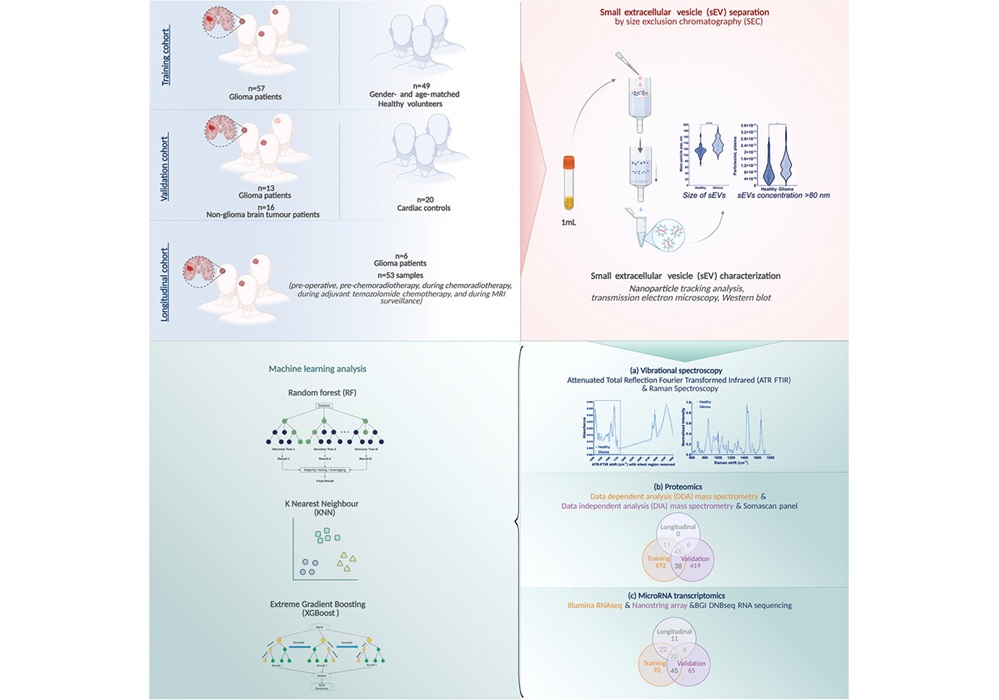
- Blood Test Enables Early Detection and Classification of Glioma
- Multi-Biomarker Blood Test Detects Early-Stage Cancers Across Types
- New Sample-to-Answer PCR System Supports High-Throughput Infectious Disease Testing
- Prognostic Biomarker Identified in Diffuse Large B-Cell Lymphoma
- Routine Blood Test Parameters Link Anemia to Cancer Risk and Mortality
- Prognostic Tool Guides Personalized Treatment in Rare Blood Cancer
- New Platelet Function Assay Enables Monitoring of Antiplatelet Therapy
- Open Multi-Omics Platform Identifies Prognostic Subtypes in Blood Cancers
- Study Finds Influenza Often Undiagnosed in Winter Deaths
- Combined Screening Approach Identifies Early Leprosy Cases
- Antibody Blood Test Identifies Active TB and Distinguishes Latent Infection
- FDA Approval Expands Use of PD-L1 Companion Diagnostic in Esophageal and GEJ Carcinomas
- Study Identifies Inflammatory Pathway Driving Immunotherapy Resistance in Bladder Cancer
- Label-Free Microscopy Methodd Enables Faster, Quantitative Detection of Malaria
- Gut Microbiome Test Predicts Melanoma Recurrence After Surgery
- Rapid Blood-Culture Susceptibility Panel Expands Coverage for Gram-Negative Infections
- Antibiotic Resistance Genes Found in Newborns Within Hours of Birth
- mNGS CSF Test Identifies CNS Pathogens Missed by Standard Panels
- Integrated System Streamlines Pre-Analytical Workflow for Molecular Testing
- Noninvasive Sputum Test Detects Early Lung Cancer
- New AI Tool Enables Rapid Treatment Selection in Pediatric Leukemia
- Breakthrough Mass Spectrometry Design Could Enable Ultra-Low Abundance Detection
- Rapid Biosensor Detects Drug Sensitivity in Breast Tumors
- Seegene Showcases Real-Time PCR Data Analytics Platform at ESCMID
- Roche Affiliate Expands MRD Portfolio with SAGA Acquisition
- Partnership Expands Ultrasensitive WGS Assay for for Hematologic Malignancies and MRD Monitoring
- QIAGEN Showcases Integrated Sample-to-Insight Oncology Solutions at AACR 2026
- Beckman Coulter Gains CE Mark for Rapid Assay Distinguishing Bacterial vs Viral Infections
- Genetic Analysis Identifies BRCA-Linked Risks Across Multiple Cancers
- Study Identifies Hidden B-Cell Mutations in Autoimmune Disease
- Single-Cell Method Measures RNA and Proteins to Reveal Immune Responses
- Study Links Midlife Vitamin D to Lower Tau in Alzheimer's
- Common Metablolic Enzyme Could Predict Response to Cancer Immunotherapy
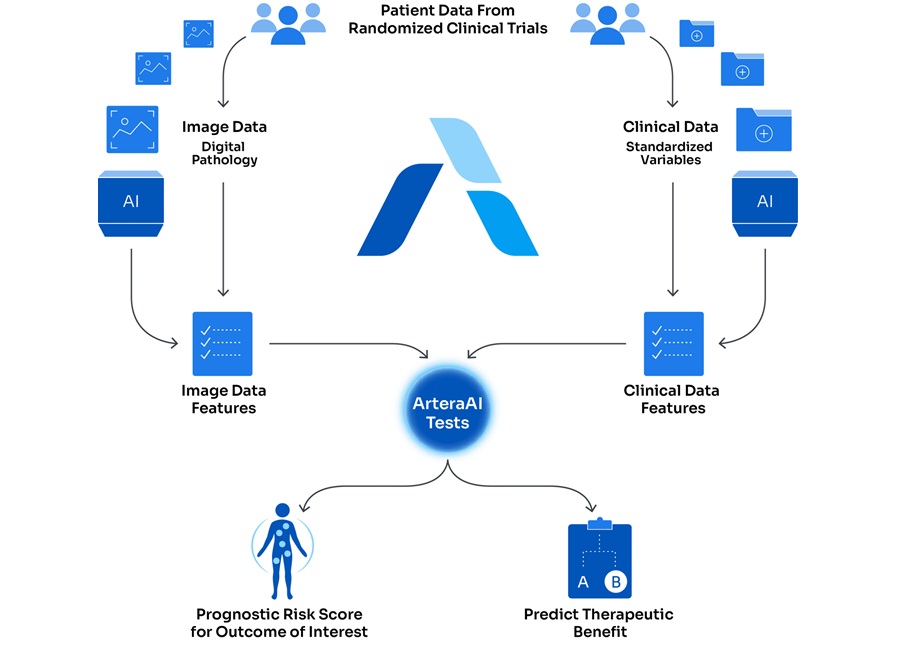
- AI-Based Assays Support Risk Stratification in Prostate and Breast Cancer
- AI Pathology Model Predicts Immunotherapy Response in Lung Cancer
- AI Precision Tests Deliver Cancer Risk Insights from Routine H&E Slides
- Study Reveals Moleclar Mechanism Driving Aggressive Skin Cancer
- Biomarker Predicts Immunotherapy Response and Prognosis in Colorectal Cancer

 Expo
Expo
- Blood Test Predicts Alzheimer Disease Risk Before Imaging Changes and Symptoms
- Study Finds ApoB Testing More Effective Than LDL for Guiding Lipid Therapy
- AI-Enabled POC Test Quantifies Multiple Cardiac Biomarkers
- Next Generation Automated Analyzers Increase Throughput for Clinical Chemistry and Electrolyte Testing
- Blood Metabolite Test Detects Early Cognitive Decline
- Composite Blood Biomarkers Enable Early Detection of Common Cancers
- Machine Learning Model Uses DNA Methylation to Predict Tumor Origin in Cancers of Unknown Primary
- Blood Test Enables Early Detection and Classification of Glioma
- Multi-Biomarker Blood Test Detects Early-Stage Cancers Across Types
- New Sample-to-Answer PCR System Supports High-Throughput Infectious Disease Testing
- Prognostic Biomarker Identified in Diffuse Large B-Cell Lymphoma
- Routine Blood Test Parameters Link Anemia to Cancer Risk and Mortality
- Prognostic Tool Guides Personalized Treatment in Rare Blood Cancer
- New Platelet Function Assay Enables Monitoring of Antiplatelet Therapy
- Open Multi-Omics Platform Identifies Prognostic Subtypes in Blood Cancers
- Study Finds Influenza Often Undiagnosed in Winter Deaths
- Combined Screening Approach Identifies Early Leprosy Cases
- Antibody Blood Test Identifies Active TB and Distinguishes Latent Infection
- FDA Approval Expands Use of PD-L1 Companion Diagnostic in Esophageal and GEJ Carcinomas
- Study Identifies Inflammatory Pathway Driving Immunotherapy Resistance in Bladder Cancer
- Label-Free Microscopy Methodd Enables Faster, Quantitative Detection of Malaria
- Gut Microbiome Test Predicts Melanoma Recurrence After Surgery
- Rapid Blood-Culture Susceptibility Panel Expands Coverage for Gram-Negative Infections
- Antibiotic Resistance Genes Found in Newborns Within Hours of Birth
- mNGS CSF Test Identifies CNS Pathogens Missed by Standard Panels
- Integrated System Streamlines Pre-Analytical Workflow for Molecular Testing
- Noninvasive Sputum Test Detects Early Lung Cancer
- New AI Tool Enables Rapid Treatment Selection in Pediatric Leukemia
- Breakthrough Mass Spectrometry Design Could Enable Ultra-Low Abundance Detection
- Rapid Biosensor Detects Drug Sensitivity in Breast Tumors
- Seegene Showcases Real-Time PCR Data Analytics Platform at ESCMID
- Roche Affiliate Expands MRD Portfolio with SAGA Acquisition
- Partnership Expands Ultrasensitive WGS Assay for for Hematologic Malignancies and MRD Monitoring
- QIAGEN Showcases Integrated Sample-to-Insight Oncology Solutions at AACR 2026
- Beckman Coulter Gains CE Mark for Rapid Assay Distinguishing Bacterial vs Viral Infections
- Genetic Analysis Identifies BRCA-Linked Risks Across Multiple Cancers
- Study Identifies Hidden B-Cell Mutations in Autoimmune Disease
- Single-Cell Method Measures RNA and Proteins to Reveal Immune Responses
- Study Links Midlife Vitamin D to Lower Tau in Alzheimer's
- Common Metablolic Enzyme Could Predict Response to Cancer Immunotherapy
- AI-Based Assays Support Risk Stratification in Prostate and Breast Cancer
- AI Pathology Model Predicts Immunotherapy Response in Lung Cancer
- AI Precision Tests Deliver Cancer Risk Insights from Routine H&E Slides
- Study Reveals Moleclar Mechanism Driving Aggressive Skin Cancer
- Biomarker Predicts Immunotherapy Response and Prognosis in Colorectal Cancer