Expo
view channel
view channel
view channel
view channel
view channel
view channel
view channel
view channel
view channel
Clinical Chem.Molecular DiagnosticsHematologyImmunologyMicrobiology
TechnologyIndustry
Events
Webinars

- Study Shows Dual Biomarkers Improve Accuracy of Alzheimer’s Detection
- Blood-Based Screening Test Targets Early Detection of Colorectal Cancer
- Automated NfL Assay Supports Monitoring of Neurological Disorders
- CSF Biomarker Improves Diagnosis of Parkinson’s Disease and Lewy Body Dementia
- Simple Urine Home Test Kit Could Detect Early-Stage Breast Cancer
- New AI Tool Improves Detection of Genetic Causes in Rare Disorders
- New Molecular Test Boosts Accuracy of Bile Duct Cancer Diagnosis
- Adaptive PCR Platform Improves Consistency in Small-Batch NGS Workflows
- First IVDR‑Certified IGH Clonality Assay Supports Diagnosis of B-Cell Malignancies
- Portable Test Uses CRISPR to Rapidly Identify STIs and Resistance Markers
- New Guidelines Aim to Improve AL Amyloidosis Diagnosis
- Fast and Easy Test Could Revolutionize Blood Transfusions
- Automated Hemostasis System Helps Labs of All Sizes Optimize Workflow
- High-Sensitivity Blood Test Improves Assessment of Clotting Risk in Heart Disease Patients
- AI Algorithm Effectively Distinguishes Alpha Thalassemia Subtypes
- Study Identifies Inflammatory Pathway Driving Immunotherapy Resistance in Bladder Cancer
- Microfluidic Chip Detects Cancer Recurrence from Immune Response Signals
- Cancer Mutation ‘Fingerprints’ to Improve Prediction of Immunotherapy Response
- Immune Signature Identified in Treatment-Resistant Myasthenia Gravis
- New Biomarker Predicts Chemotherapy Response in Triple-Negative Breast Cancer
- Genomic Analysis Links Emerging Streptococcal Strains to Specific Infections
- Rapid Urine Test Speeds Antibiotic Selection for UTIs
- WHO Endorses Rapid Point-of-Care Testing to Improve TB Detection
- Breath Analysis Approach Offers Rapid Detection of Bacterial Infection
- Study Highlights Accuracy Gaps in Consumer Gut Microbiome Kits
- Breakthrough Mass Spectrometry Design Could Enable Ultra-Low Abundance Detection
- Rapid Biosensor Detects Drug Sensitivity in Breast Tumors
- Online Tool Supports Family Screening for Inherited Cancer Risk
- Portable Breath Sensor Detects Pneumonia Biomarkers in Minutes
- New Electronic Pipette Enhances Workflows with Touchscreen Control
- Lunit and CellCarta Collaborate to Expand AI Pathology in CDx Development
- Integrated DNA Technologies Expands into Clinical Diagnostics
- Co-Diagnostics Agreement Expands Commercial and Distribution Reach in South Asia
- Automated MSI Test Gains IVDR Certification to Guide CRC Therapy
- New Partnership Brings Alzheimer’s Blood Biomarker Test to Community Screening Network
- Study Reveals Diagnostic and Therapeutic Target in Rare Pancreatic Tumors
- Large-Scale Study Maps DNA Damage Signatures Across Multiple Cancers
- Study Identifies Distinct Immune Signatures to Early Depression and Psychosis
- Genetic Mutation Behind Aggressive Adult Leukemia Offers Treatment Clues
- Disease Gene Discovery Advances Diagnosis of Rare Movement Disorders
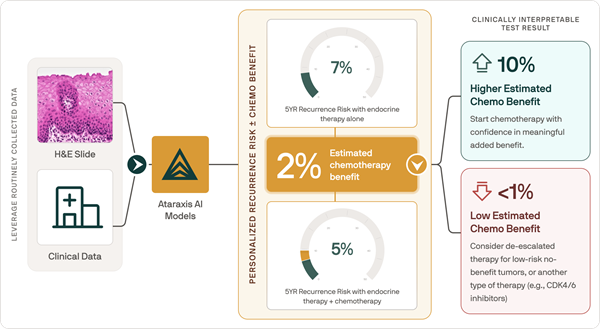
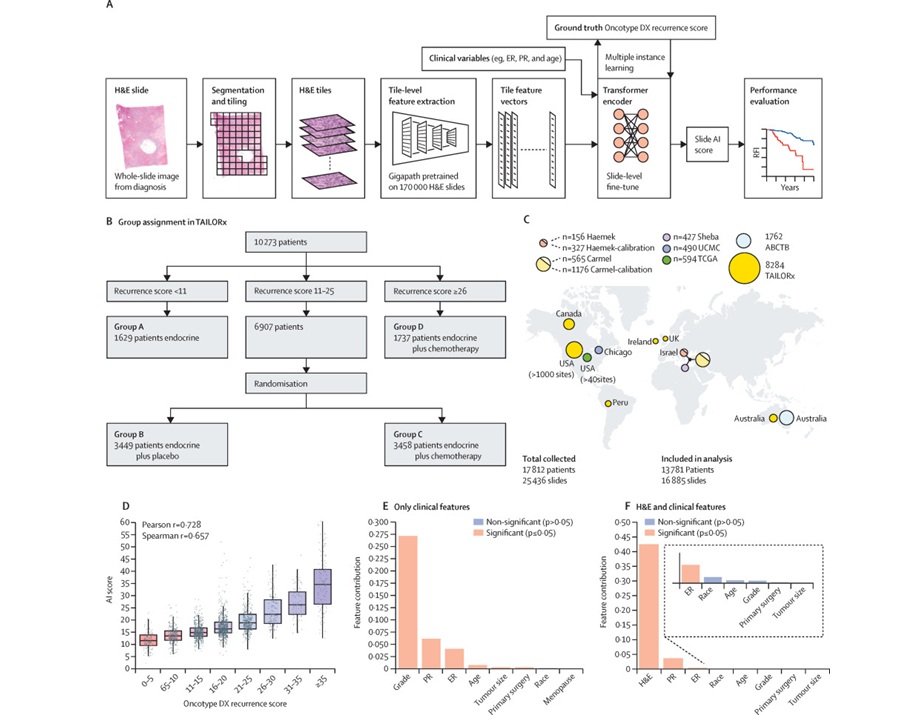
- AI-Based Pathology Model Guides Chemotherapy Decisions in Breast Cancer
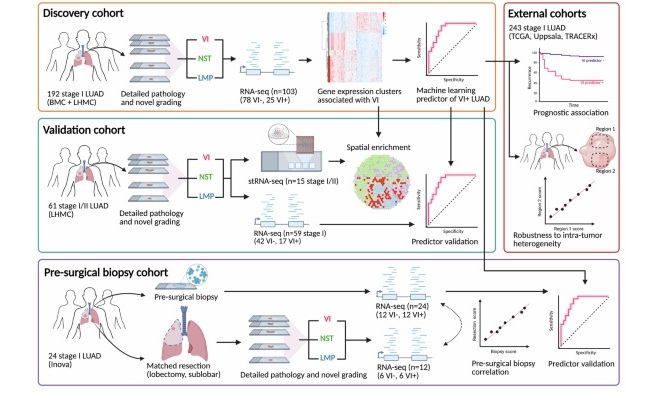
- Biopsy-Based Gene Test Predicts Recurrence Risk in Lung Adenocarcinoma
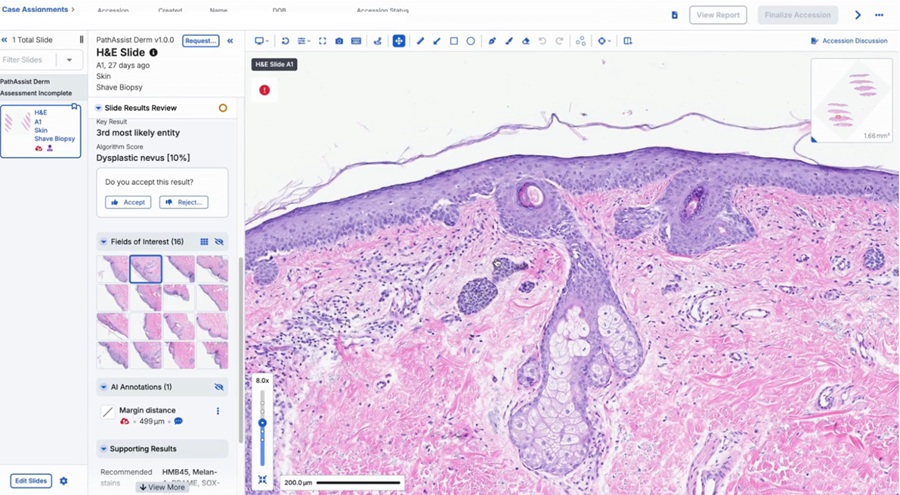
- AI-Powered Tool to Transform Dermatopathology Workflow
- New Chromogenic Culture Media Enable Rapid Detection of Candida Infections
- Novel mcPCR Technology to Transform Testing of Clinical Samples

 Expo
Expo
- Study Shows Dual Biomarkers Improve Accuracy of Alzheimer’s Detection
- Blood-Based Screening Test Targets Early Detection of Colorectal Cancer
- Automated NfL Assay Supports Monitoring of Neurological Disorders
- CSF Biomarker Improves Diagnosis of Parkinson’s Disease and Lewy Body Dementia
- Simple Urine Home Test Kit Could Detect Early-Stage Breast Cancer
- New AI Tool Improves Detection of Genetic Causes in Rare Disorders
- New Molecular Test Boosts Accuracy of Bile Duct Cancer Diagnosis
- Adaptive PCR Platform Improves Consistency in Small-Batch NGS Workflows
- First IVDR‑Certified IGH Clonality Assay Supports Diagnosis of B-Cell Malignancies
- Portable Test Uses CRISPR to Rapidly Identify STIs and Resistance Markers
- New Guidelines Aim to Improve AL Amyloidosis Diagnosis
- Fast and Easy Test Could Revolutionize Blood Transfusions
- Automated Hemostasis System Helps Labs of All Sizes Optimize Workflow
- High-Sensitivity Blood Test Improves Assessment of Clotting Risk in Heart Disease Patients
- AI Algorithm Effectively Distinguishes Alpha Thalassemia Subtypes
- Study Identifies Inflammatory Pathway Driving Immunotherapy Resistance in Bladder Cancer
- Microfluidic Chip Detects Cancer Recurrence from Immune Response Signals
- Cancer Mutation ‘Fingerprints’ to Improve Prediction of Immunotherapy Response
- Immune Signature Identified in Treatment-Resistant Myasthenia Gravis
- New Biomarker Predicts Chemotherapy Response in Triple-Negative Breast Cancer
- Genomic Analysis Links Emerging Streptococcal Strains to Specific Infections
- Rapid Urine Test Speeds Antibiotic Selection for UTIs
- WHO Endorses Rapid Point-of-Care Testing to Improve TB Detection
- Breath Analysis Approach Offers Rapid Detection of Bacterial Infection
- Study Highlights Accuracy Gaps in Consumer Gut Microbiome Kits
- Breakthrough Mass Spectrometry Design Could Enable Ultra-Low Abundance Detection
- Rapid Biosensor Detects Drug Sensitivity in Breast Tumors
- Online Tool Supports Family Screening for Inherited Cancer Risk
- Portable Breath Sensor Detects Pneumonia Biomarkers in Minutes
- New Electronic Pipette Enhances Workflows with Touchscreen Control
- Lunit and CellCarta Collaborate to Expand AI Pathology in CDx Development
- Integrated DNA Technologies Expands into Clinical Diagnostics
- Co-Diagnostics Agreement Expands Commercial and Distribution Reach in South Asia
- Automated MSI Test Gains IVDR Certification to Guide CRC Therapy
- New Partnership Brings Alzheimer’s Blood Biomarker Test to Community Screening Network
- Study Reveals Diagnostic and Therapeutic Target in Rare Pancreatic Tumors
- Large-Scale Study Maps DNA Damage Signatures Across Multiple Cancers
- Study Identifies Distinct Immune Signatures to Early Depression and Psychosis
- Genetic Mutation Behind Aggressive Adult Leukemia Offers Treatment Clues
- Disease Gene Discovery Advances Diagnosis of Rare Movement Disorders
- AI-Based Pathology Model Guides Chemotherapy Decisions in Breast Cancer
- Biopsy-Based Gene Test Predicts Recurrence Risk in Lung Adenocarcinoma
- AI-Powered Tool to Transform Dermatopathology Workflow
- New Chromogenic Culture Media Enable Rapid Detection of Candida Infections
- Novel mcPCR Technology to Transform Testing of Clinical Samples